![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-104_TAAGGCGA-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 734874 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
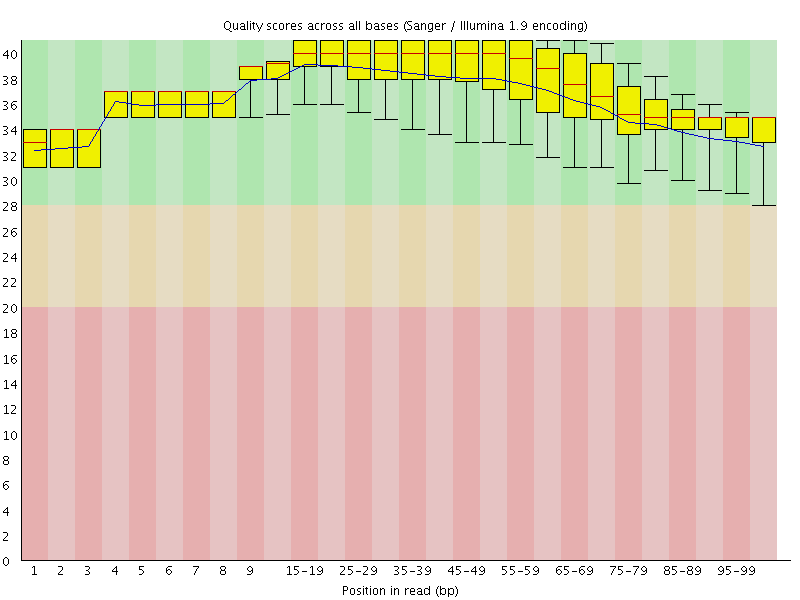
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
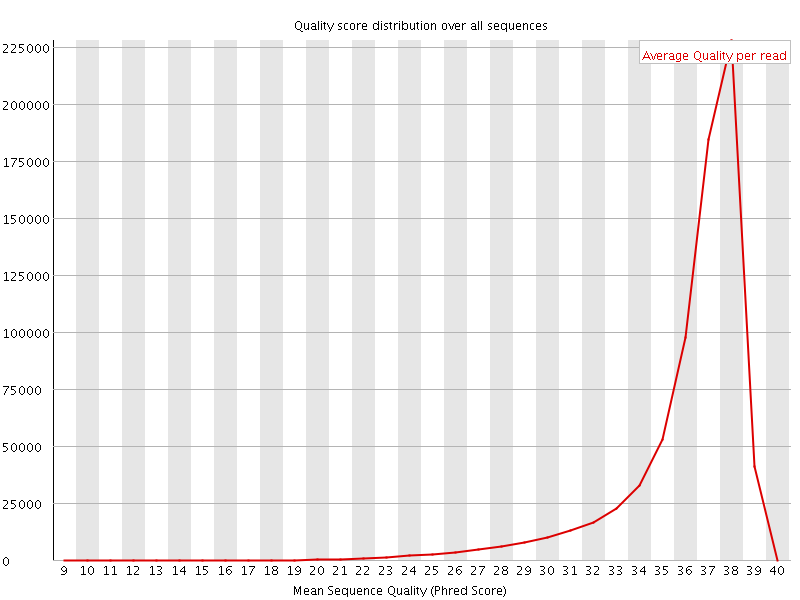
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
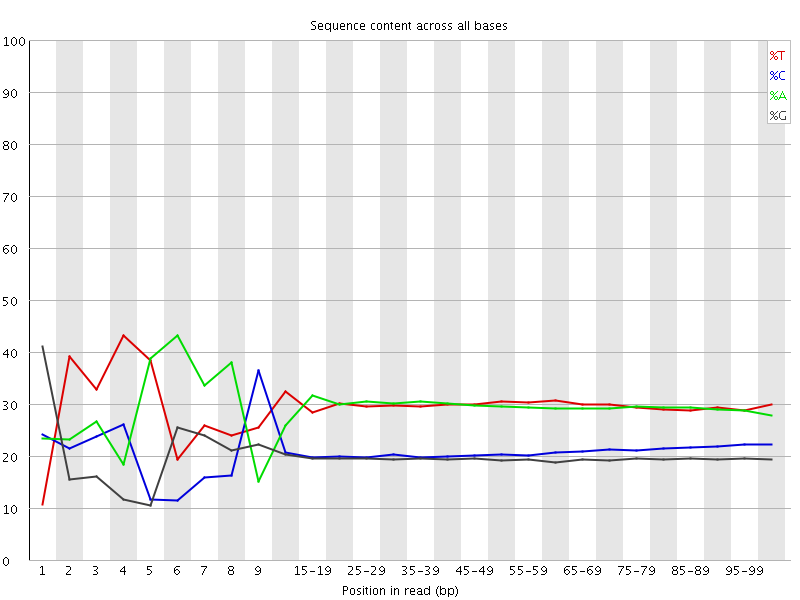
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
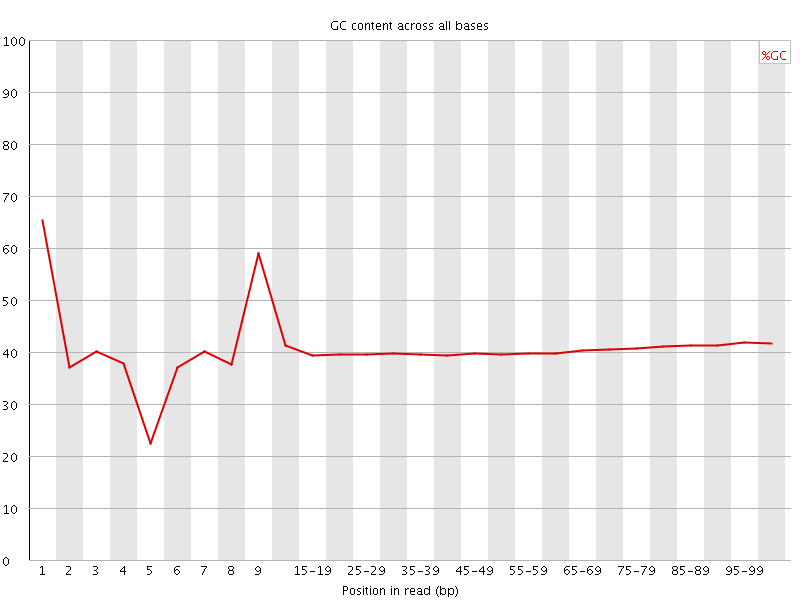
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
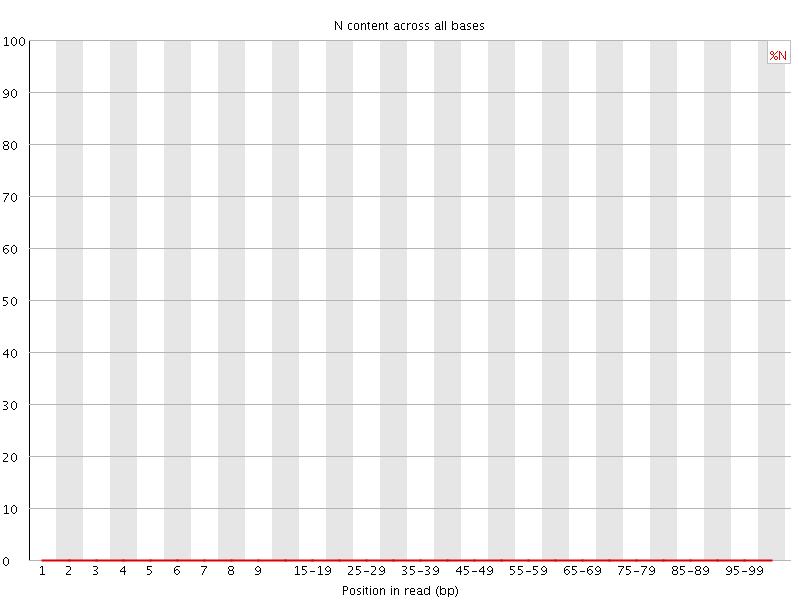
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
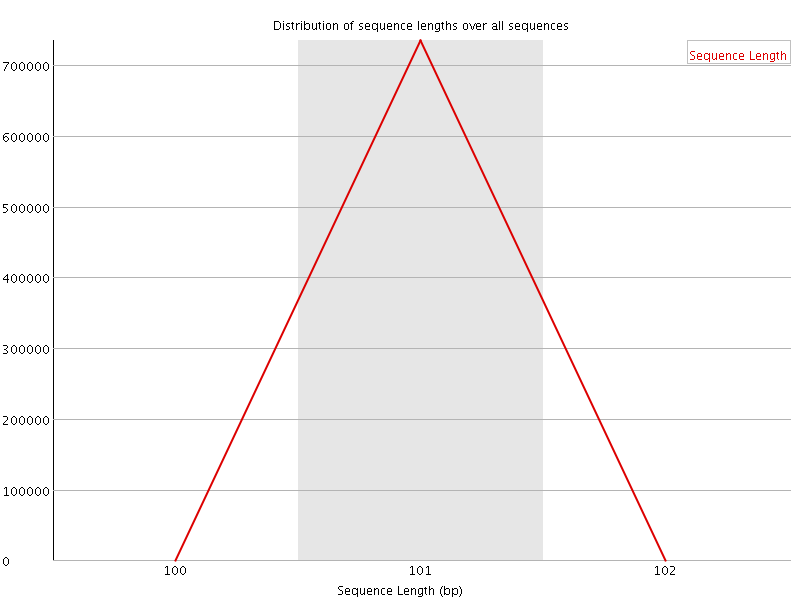
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
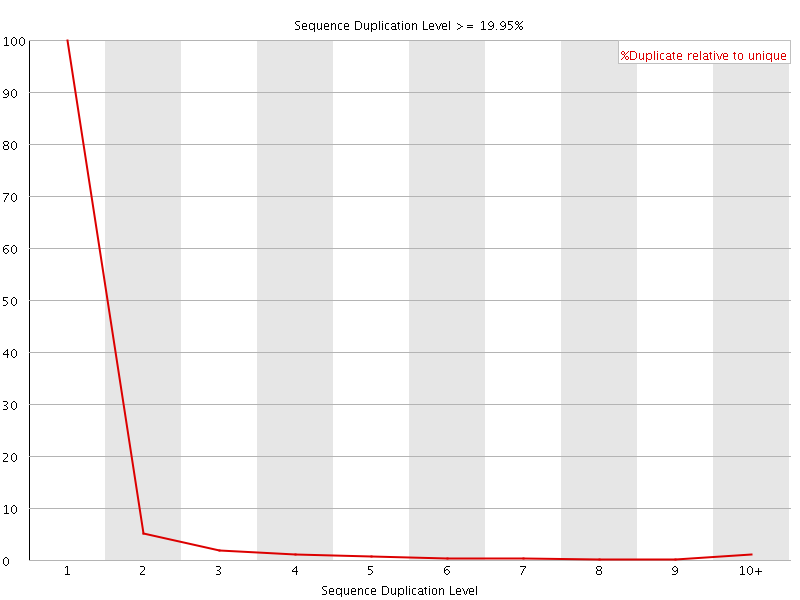
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
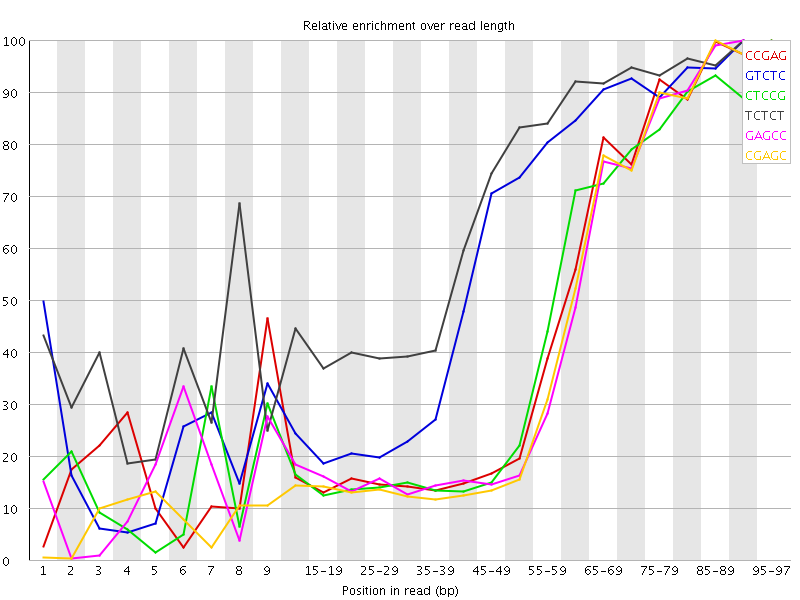
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 112705 | 3.171375 | 7.149321 | 95-97 |
| GTCTC | 171975 | 3.1605482 | 5.346733 | 90-94 |
| CTCCG | 116825 | 3.0787976 | 7.0706816 | 95-97 |
| TCTCT | 250270 | 3.0339355 | 4.4379735 | 90-94 |
| GAGCC | 106900 | 3.0080292 | 7.026705 | 90-94 |
| CGAGC | 102670 | 2.8890026 | 6.9504685 | 85-89 |
| AGAAG | 204465 | 2.8539243 | 5.001236 | 5 |
| TCTCC | 159620 | 2.7748096 | 5.220348 | 95-97 |
| GCCCA | 101155 | 2.692406 | 6.5846663 | 90-94 |
| AGCCC | 100200 | 2.6669872 | 6.4994864 | 90-94 |
| ACACA | 208645 | 2.6057281 | 7.762803 | 6 |
| ATCTC | 203215 | 2.4880621 | 5.6852417 | 95-97 |
| CCCAC | 98280 | 2.474384 | 6.2382536 | 90-94 |
| CAAAG | 181295 | 2.393636 | 5.332449 | 4 |
| CCACG | 88170 | 2.346789 | 6.453025 | 90-94 |
| TACAC | 188400 | 2.3296688 | 10.246245 | 5 |
| TCCGA | 125225 | 2.32432 | 5.033901 | 95-97 |
| CTTTG | 176105 | 2.2569425 | 8.947892 | 9 |
| CACAT | 180285 | 2.2293222 | 5.5398073 | 7 |
| GAGAC | 110525 | 2.1904051 | 5.426067 | 95-97 |
| CGAGA | 106800 | 2.1165824 | 5.3427863 | 95-97 |
| ACGAG | 99570 | 1.9732969 | 5.166616 | 95-97 |
| GGCGA | 65755 | 1.9560698 | 6.6206 | 95-97 |
| AGGCG | 65705 | 1.9545823 | 6.5580964 | 95-97 |
| GCTTT | 147310 | 1.8879089 | 5.8101525 | 1 |
| TGGAG | 86110 | 1.7863265 | 5.0391164 | 5 |
| GAGAG | 83695 | 1.7535357 | 5.7598023 | 7 |
| AGACT | 132635 | 1.7338942 | 6.1735196 | 6 |
| GCCAA | 88265 | 1.6546315 | 5.3628035 | 1 |
| TTGAG | 120485 | 1.6486967 | 5.9446354 | 9 |
| TGGCT | 84180 | 1.6355249 | 5.3129625 | 6 |
| TATAC | 186205 | 1.6056701 | 5.6526365 | 5 |
| GAGTC | 77425 | 1.519278 | 7.82055 | 9 |
| GACTC | 81450 | 1.5118057 | 5.714629 | 7 |
| GTCTT | 114675 | 1.4696623 | 6.6241956 | 1 |
| AAGAC | 109365 | 1.4439452 | 6.049417 | 5 |
| CATGA | 108520 | 1.4186467 | 5.41926 | 9 |
| TGAGT | 101280 | 1.3858987 | 5.546557 | 8 |
| GTGTT | 101690 | 1.3777746 | 6.661393 | 1 |
| TATGA | 150255 | 1.369762 | 6.5593295 | 4 |
| GACTT | 105715 | 1.3683375 | 7.2798767 | 7 |
| CCATG | 73705 | 1.3680496 | 5.9216156 | 9 |
| ATGAG | 94470 | 1.3055981 | 5.280211 | 7 |
| GTATG | 94395 | 1.2916855 | 6.030885 | 3 |
| TCATA | 148830 | 1.2833806 | 6.568532 | 2 |
| GATTG | 93130 | 1.2743754 | 6.1370397 | 7 |
| AACAC | 101730 | 1.2704868 | 5.4012637 | 5 |
| GTGTA | 91230 | 1.2483761 | 11.325789 | 1 |
| GTTCT | 97220 | 1.2459607 | 5.201174 | 1 |
| ACATG | 89065 | 1.1643177 | 5.805897 | 8 |
| ACCAT | 93300 | 1.1537054 | 5.0361886 | 8 |
| TGGAC | 57870 | 1.1355586 | 7.849092 | 5 |
| ACACC | 63355 | 1.1234208 | 5.467995 | 6 |
| ACACT | 88900 | 1.0992969 | 5.8635626 | 6 |
| GTCCA | 59025 | 1.095572 | 7.8568115 | 1 |
| GGACT | 55030 | 1.0798305 | 7.5541573 | 6 |
| GAGTA | 74520 | 1.0298843 | 5.5082617 | 1 |
| TGTAT | 113545 | 1.0248877 | 5.0024376 | 2 |
| CATAC | 76205 | 0.94231623 | 5.557793 | 3 |
| TGCCG | 31425 | 0.87553155 | 5.1064305 | 95-97 |
| GTATA | 83985 | 0.76562816 | 6.4269915 | 1 |