![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-102_TAAGGCGA-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1697969 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
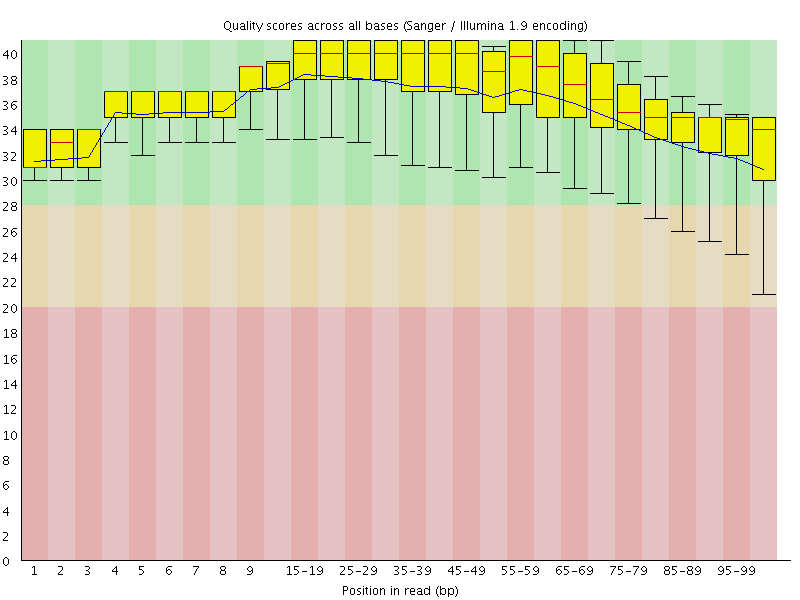
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
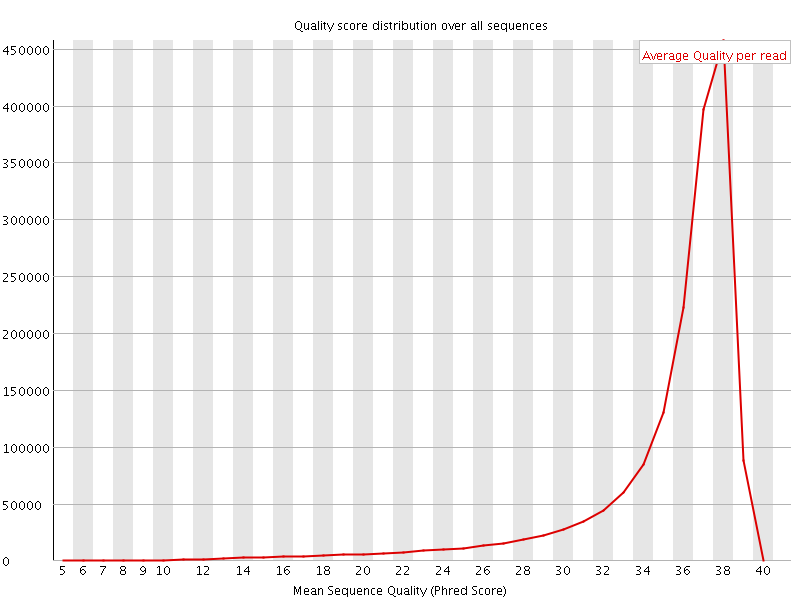
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
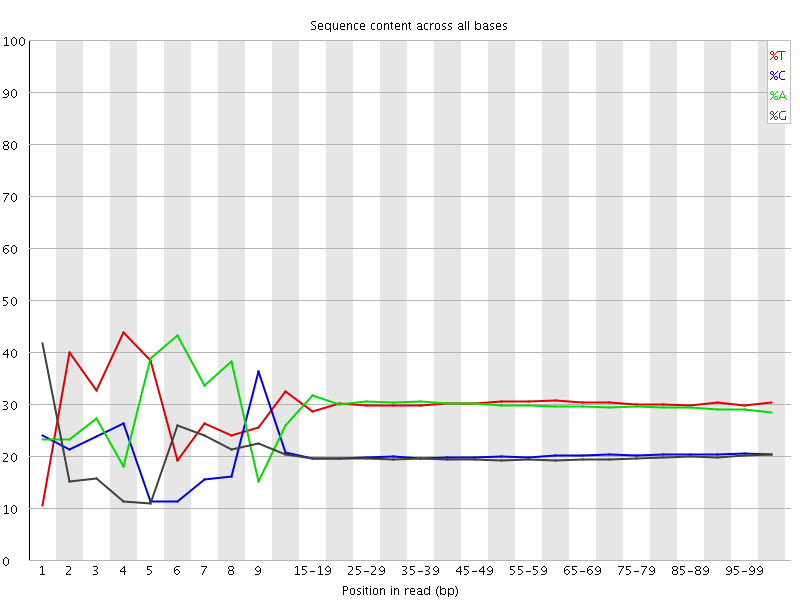
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
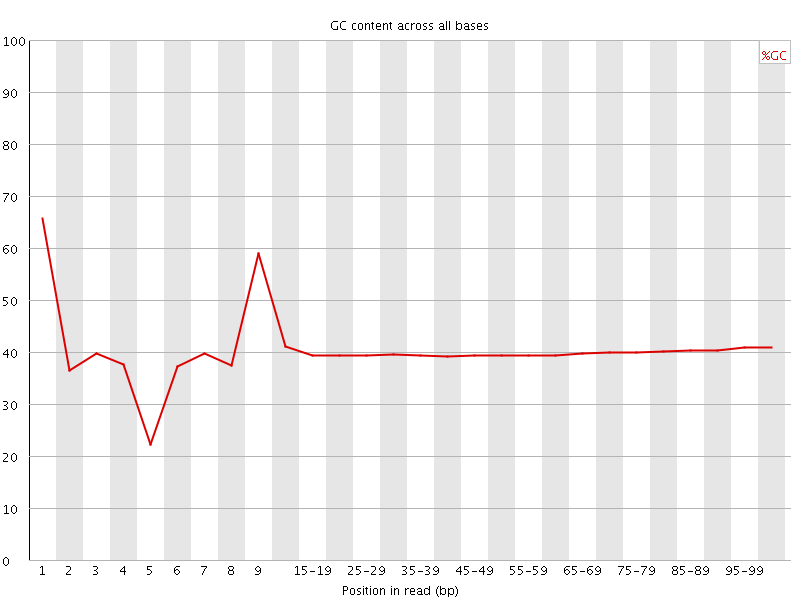
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
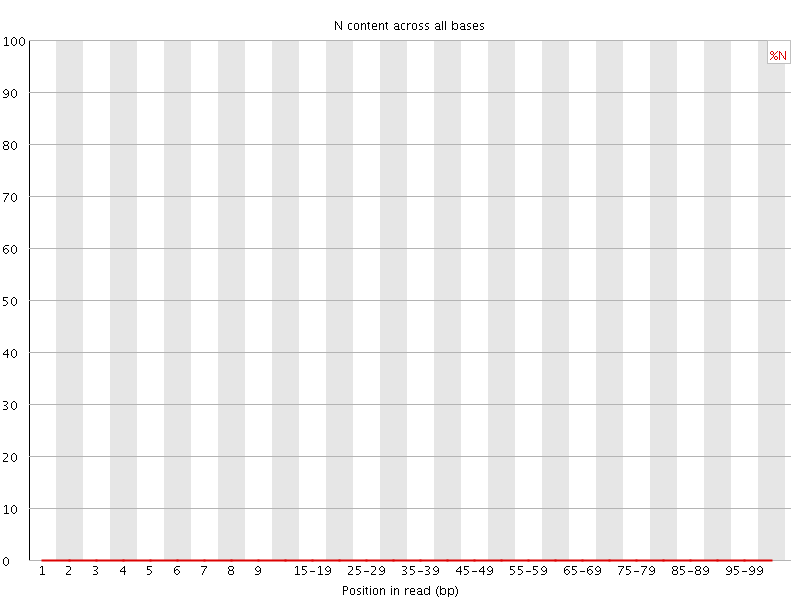
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
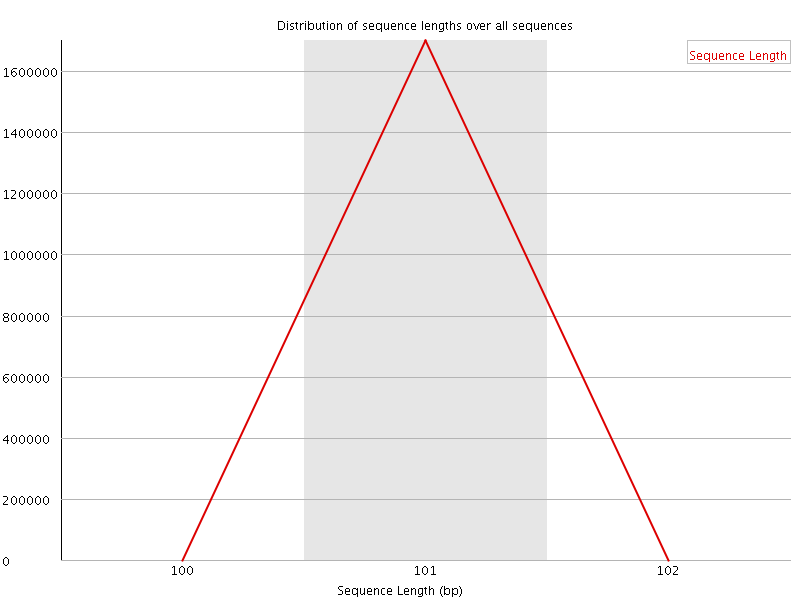
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
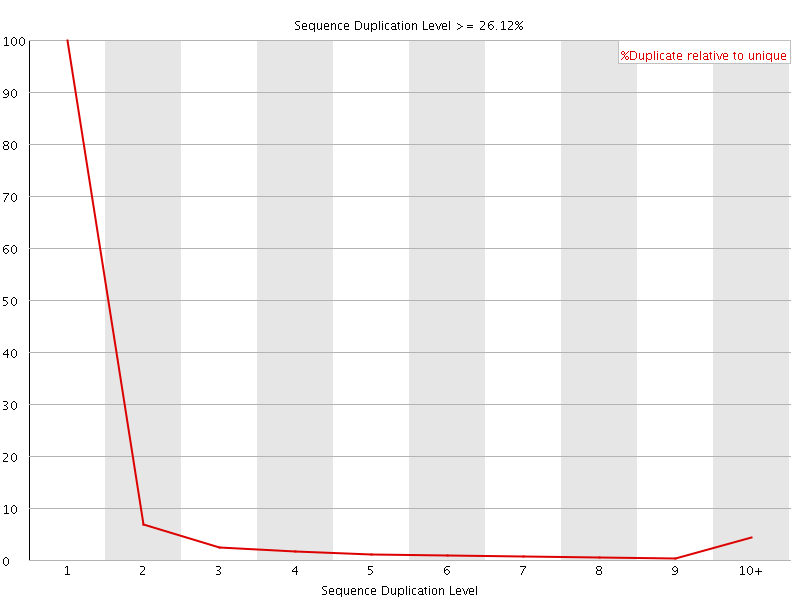
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
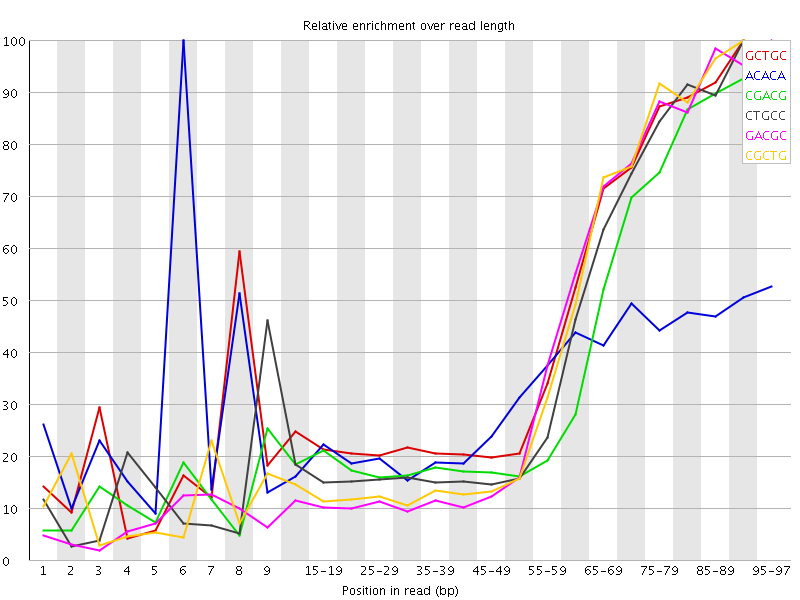
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 206450 | 2.5625136 | 5.6212397 | 90-94 |
| ACACA | 401115 | 2.2677023 | 6.95824 | 6 |
| CGACG | 169835 | 2.1429904 | 5.520289 | 95-97 |
| CTGCC | 175925 | 2.141553 | 5.2059293 | 90-94 |
| GACGC | 169560 | 2.1395202 | 5.279567 | 95-97 |
| CGCTG | 170995 | 2.1224363 | 5.13364 | 90-94 |
| GCCGA | 168025 | 2.1201515 | 5.452968 | 95-97 |
| CTTTG | 374380 | 2.0542562 | 7.860464 | 9 |
| CCGAC | 161625 | 2.0000992 | 5.2038445 | 95-97 |
| TACAC | 351605 | 1.955377 | 8.772927 | 5 |
| TGCCG | 153590 | 1.9064009 | 5.0610123 | 95-97 |
| GAGAG | 205360 | 1.8033122 | 6.0471134 | 7 |
| GCTTT | 318150 | 1.7457174 | 5.1398354 | 1 |
| TGGAG | 191980 | 1.6583238 | 5.0952497 | 5 |
| TTGAG | 258765 | 1.4717674 | 5.053065 | 9 |
| GAGTC | 165435 | 1.4014924 | 6.8279467 | 9 |
| CCATG | 165535 | 1.3753182 | 5.668957 | 9 |
| AAGAC | 237475 | 1.3689419 | 5.3437524 | 5 |
| TATAC | 360295 | 1.3452531 | 5.641861 | 5 |
| GTGTA | 236095 | 1.3428283 | 10.398837 | 1 |
| GTCTT | 237980 | 1.3058175 | 6.4041657 | 1 |
| GACTT | 223940 | 1.2491523 | 6.0973673 | 7 |
| AACAC | 219860 | 1.2429776 | 5.0817237 | 5 |
| TATGA | 324180 | 1.2341897 | 5.6141996 | 4 |
| GTGTT | 220555 | 1.2339822 | 6.1391683 | 1 |
| TCATA | 323050 | 1.2061894 | 5.721469 | 2 |
| GATTG | 211180 | 1.2011201 | 5.626891 | 7 |
| GTTCT | 215075 | 1.1801355 | 5.355437 | 1 |
| ACACC | 138025 | 1.1433069 | 5.4228735 | 6 |
| TGGAC | 127125 | 1.0769469 | 6.9490514 | 5 |
| GTCCA | 128600 | 1.0684503 | 7.210222 | 1 |
| ACACT | 189115 | 1.0517231 | 5.1133366 | 6 |
| AGACT | 177705 | 1.0076855 | 5.1450524 | 6 |
| GGACT | 117915 | 0.99892384 | 6.548507 | 6 |
| GAGTA | 166100 | 0.9603842 | 5.003745 | 1 |
| GTATG | 166525 | 0.94713765 | 5.018523 | 3 |
| GTATA | 196695 | 0.74884003 | 7.062176 | 1 |