![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-102_TAAGGCGA-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1697969 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
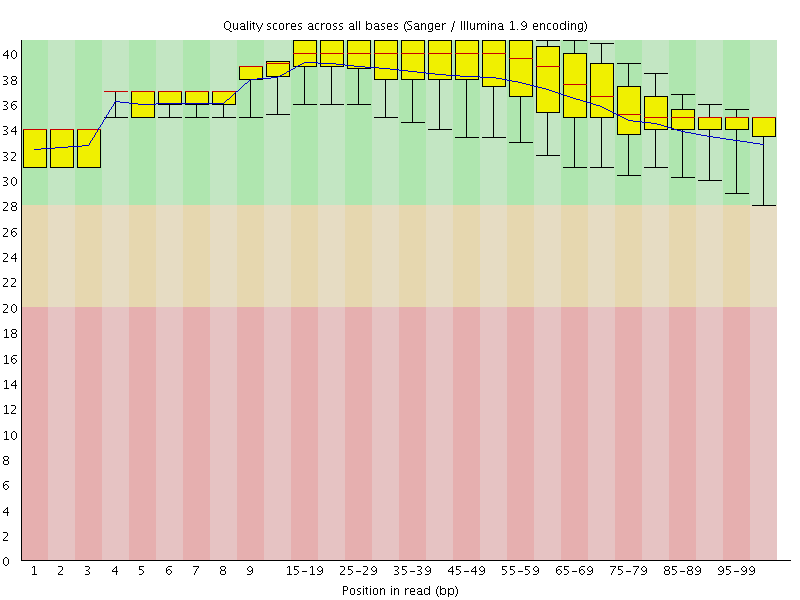
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
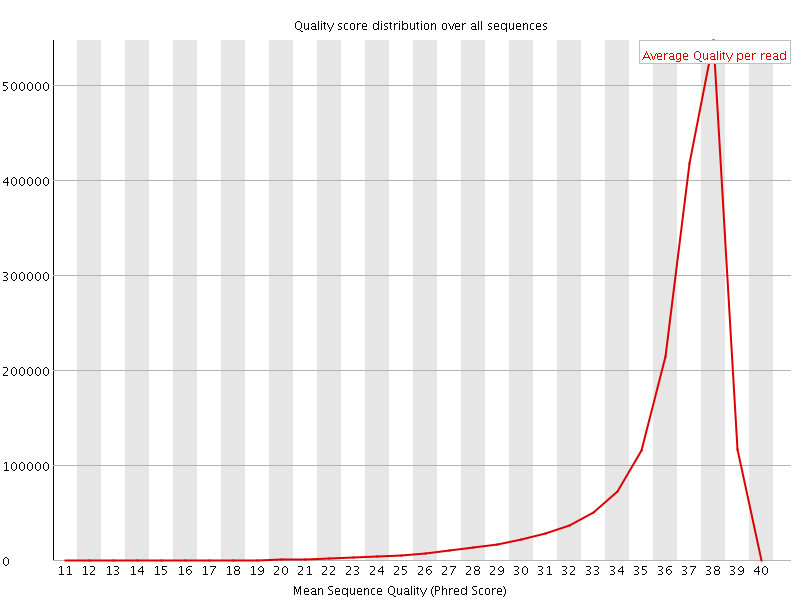
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
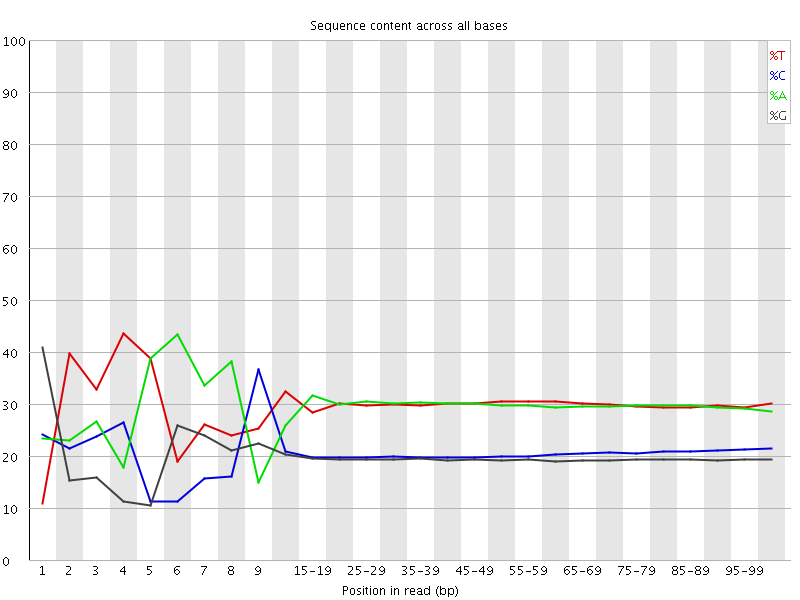
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
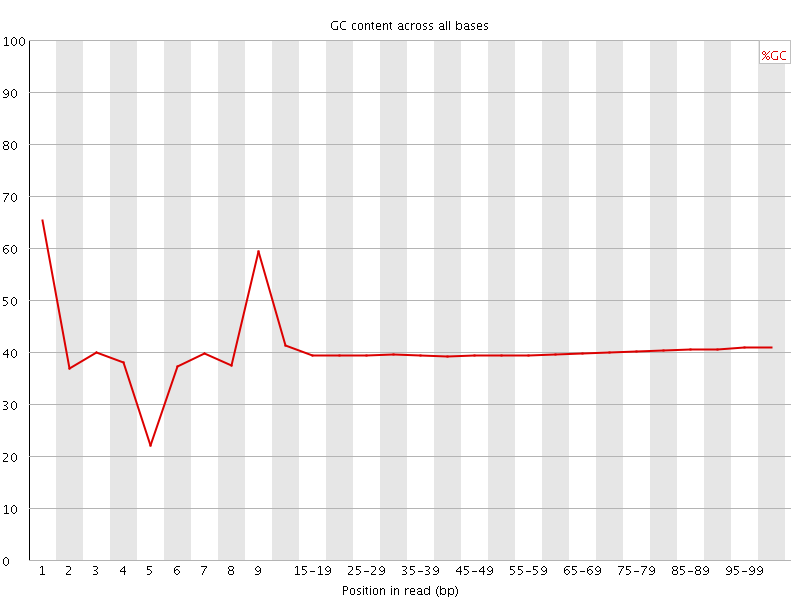
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
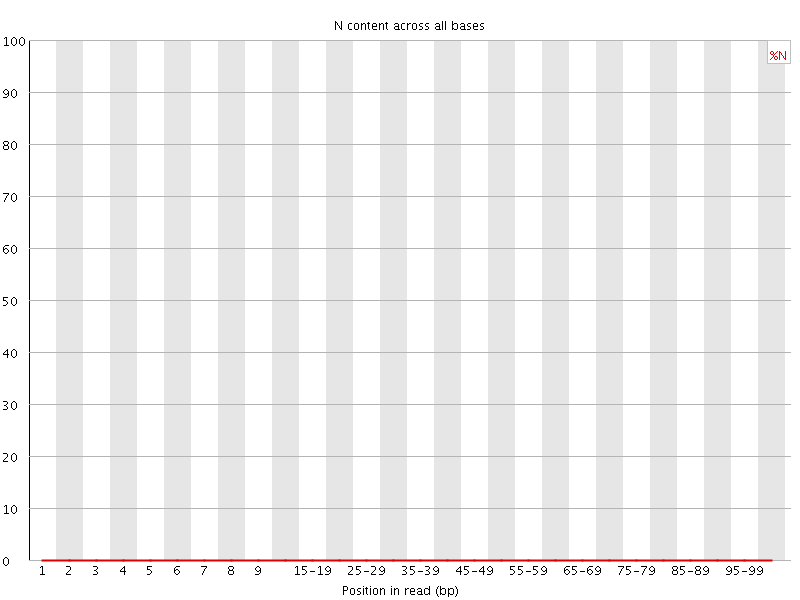
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
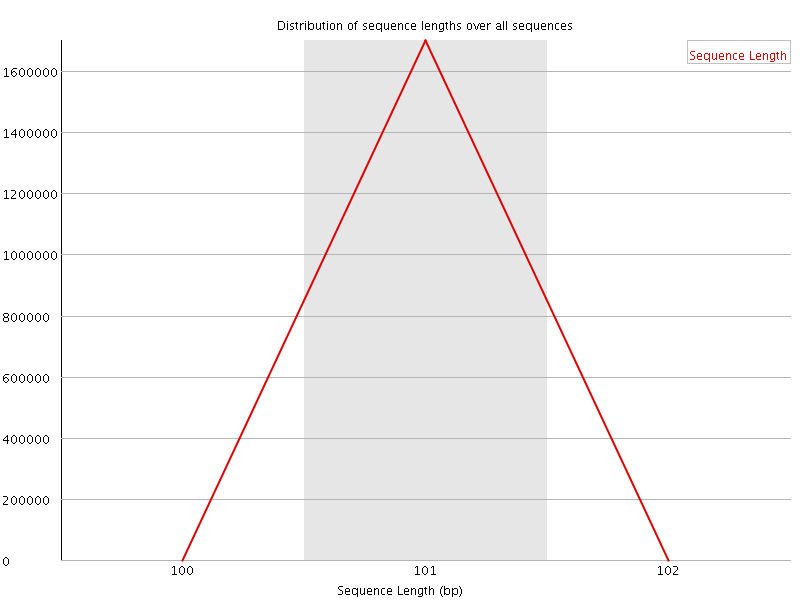
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
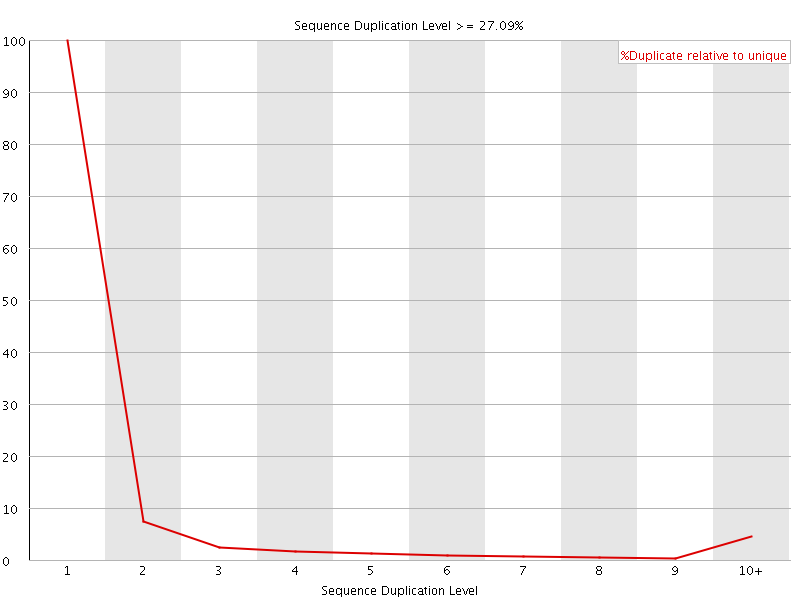
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
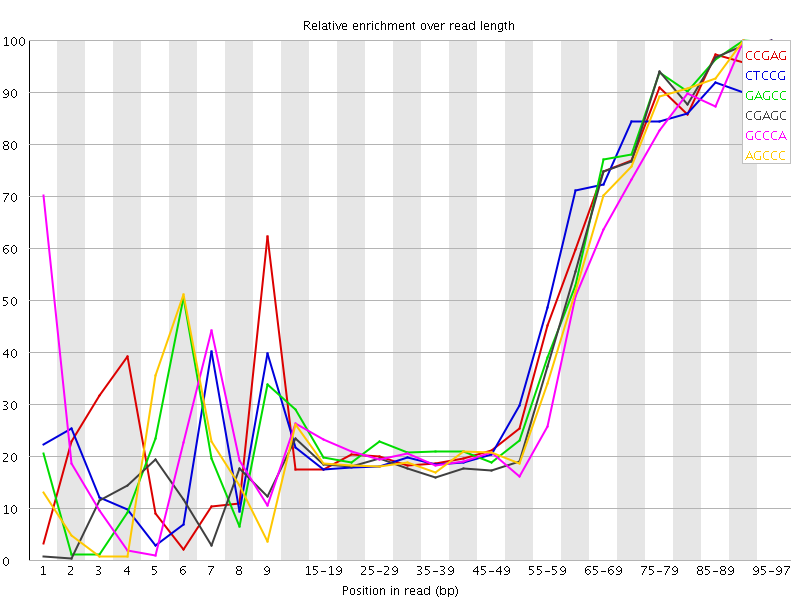
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 213305 | 2.666502 | 5.7158933 | 95-97 |
| CTCCG | 221585 | 2.628094 | 5.620498 | 95-97 |
| GAGCC | 208255 | 2.603372 | 5.50505 | 90-94 |
| CGAGC | 193895 | 2.4238594 | 5.408782 | 95-97 |
| GCCCA | 198135 | 2.3726082 | 5.36972 | 90-94 |
| AGCCC | 195485 | 2.3408754 | 5.212949 | 90-94 |
| ACACA | 404385 | 2.2111335 | 6.834512 | 6 |
| CCCAC | 187205 | 2.1473672 | 5.0435853 | 90-94 |
| CTTTG | 377445 | 2.0934212 | 7.9597487 | 9 |
| TACAC | 354745 | 1.9211972 | 8.804281 | 5 |
| GAGAG | 203175 | 1.8306384 | 6.34313 | 7 |
| GCTTT | 318280 | 1.7652748 | 5.085815 | 1 |
| TTGAG | 259510 | 1.517041 | 5.1952972 | 9 |
| AGACT | 262645 | 1.484912 | 5.5234613 | 6 |
| GAGTC | 166055 | 1.4195281 | 6.776641 | 9 |
| GACTC | 170490 | 1.3960949 | 5.010489 | 7 |
| GTCTT | 249730 | 1.3850762 | 6.333736 | 1 |
| CCATG | 169085 | 1.3845899 | 6.1142263 | 9 |
| AAGAC | 237355 | 1.3548595 | 5.648636 | 5 |
| TATAC | 364600 | 1.3502787 | 5.817769 | 5 |
| GTGTT | 221100 | 1.28017 | 6.350934 | 1 |
| TGAGT | 217855 | 1.2735345 | 5.1981316 | 8 |
| GACTT | 224500 | 1.2571399 | 6.2228103 | 7 |
| TATGA | 323035 | 1.2489134 | 5.6741376 | 4 |
| GATTG | 206600 | 1.2077402 | 5.6289473 | 7 |
| TCATA | 325035 | 1.2037518 | 5.6767197 | 2 |
| GTTCT | 213880 | 1.1862415 | 5.279458 | 1 |
| GTGTA | 195630 | 1.1436119 | 10.644353 | 1 |
| ACCAT | 207245 | 1.1223794 | 5.201694 | 8 |
| ACACC | 138865 | 1.0997583 | 5.183736 | 6 |
| TGGAC | 125700 | 1.0745517 | 6.7351937 | 5 |
| GTCCA | 130855 | 1.071535 | 7.4095836 | 1 |
| ACACT | 187860 | 1.0173959 | 5.1202946 | 6 |
| GGACT | 117295 | 1.0027012 | 6.5030885 | 6 |
| GAGTA | 165235 | 0.9752357 | 5.0829587 | 1 |
| GTATA | 197300 | 0.7627986 | 6.857923 | 1 |