![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-101_TAAGGCGA-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1584621 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
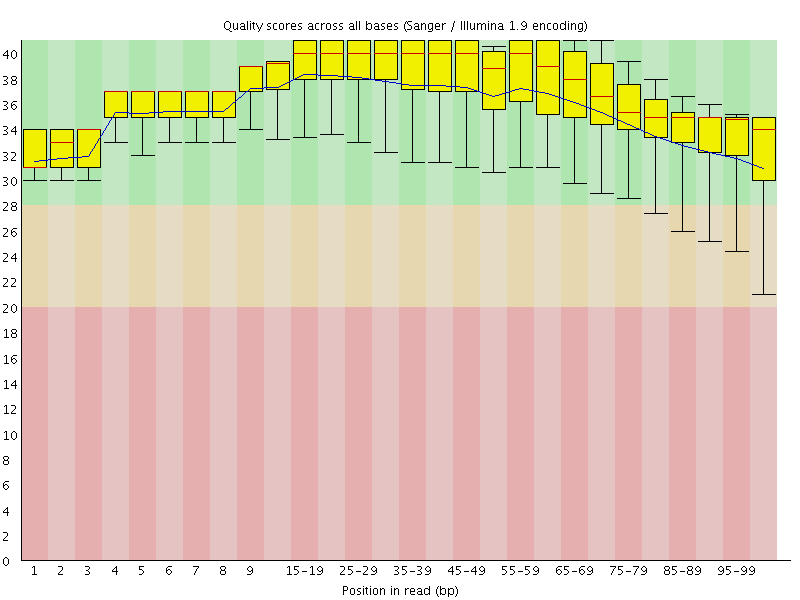
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
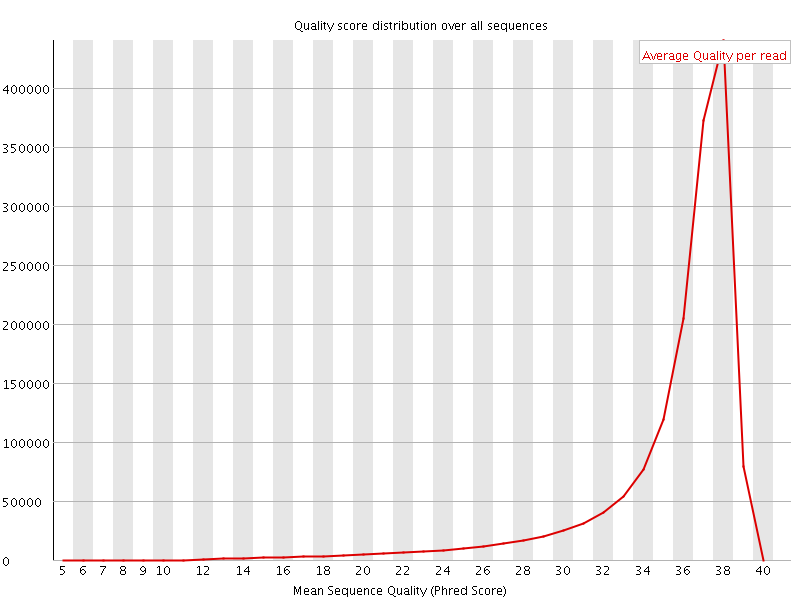
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
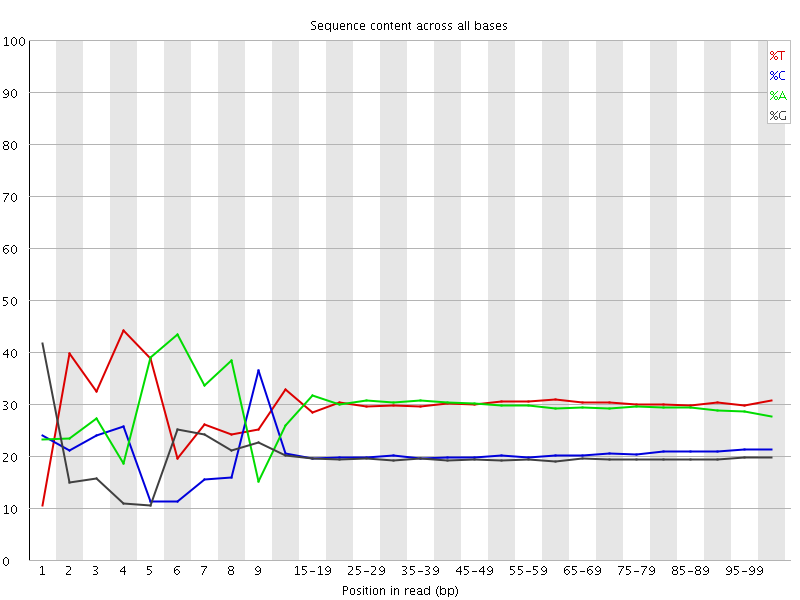
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
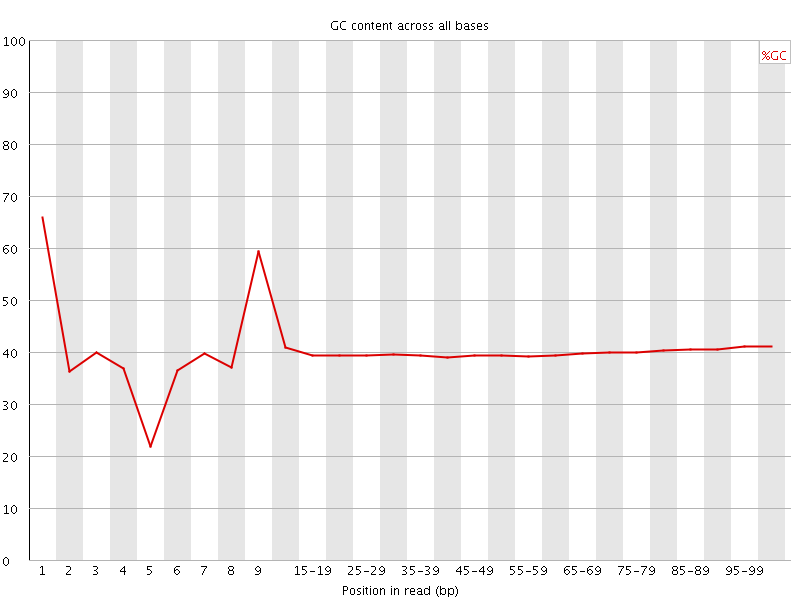
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
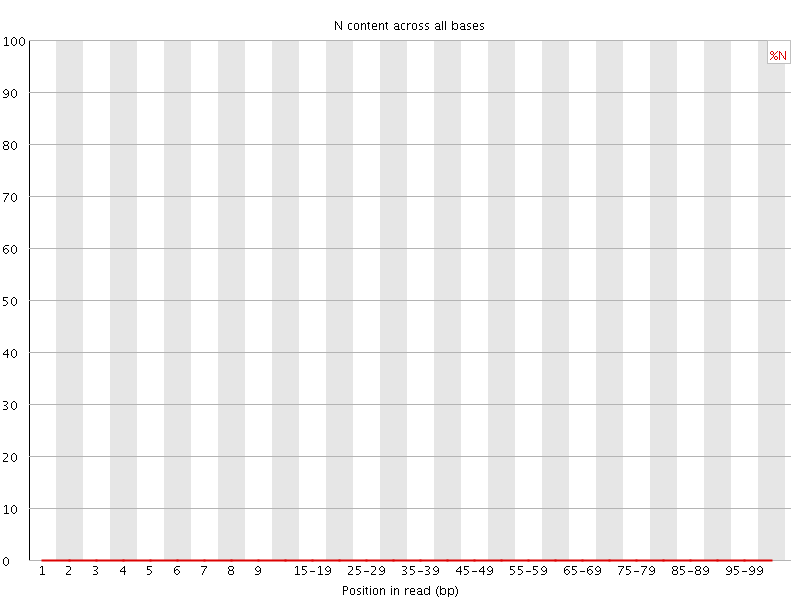
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
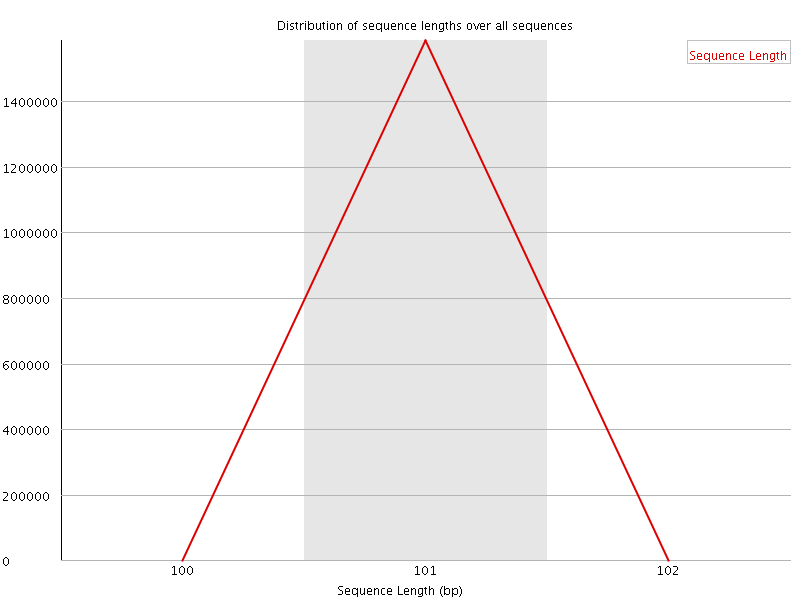
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
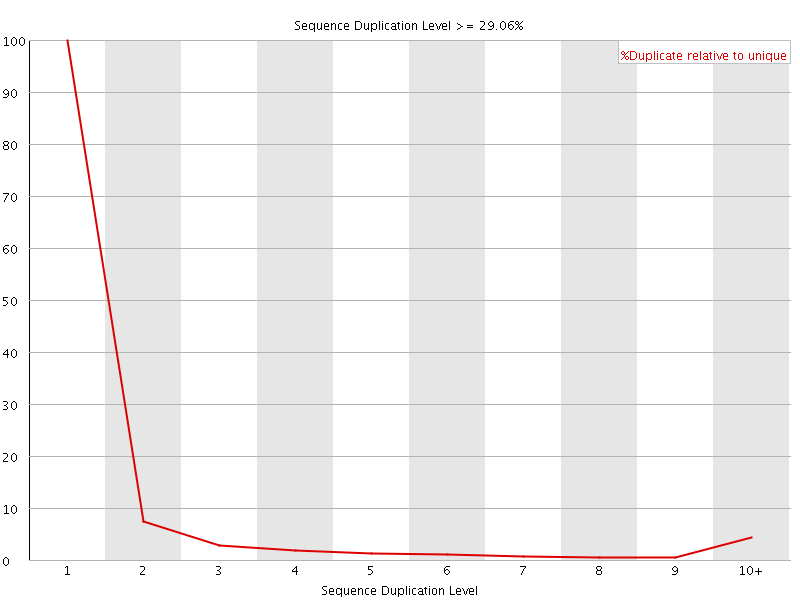
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1810 | 0.11422289620041637 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
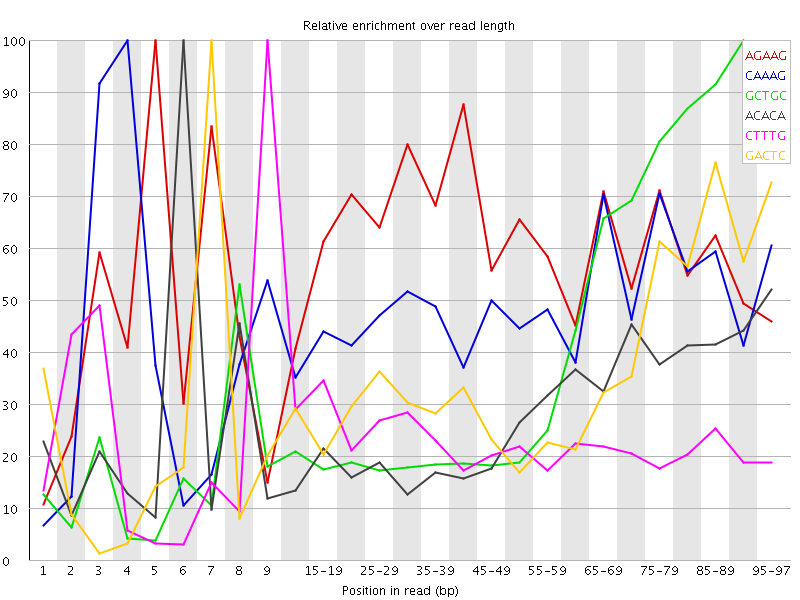
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 467840 | 3.0130396 | 4.998458 | 5 |
| CAAAG | 431130 | 2.675108 | 5.514004 | 4 |
| GCTGC | 191795 | 2.546234 | 6.039468 | 90-94 |
| ACACA | 420225 | 2.5121207 | 8.833247 | 6 |
| CTTTG | 422400 | 2.4736106 | 10.686895 | 9 |
| GACTC | 272115 | 2.4009032 | 6.6620984 | 7 |
| TACAC | 382610 | 2.24357 | 11.152925 | 5 |
| GACGC | 162575 | 2.2003407 | 5.80739 | 95-97 |
| GCCGA | 160360 | 2.1703625 | 5.8817883 | 95-97 |
| AAAGC | 349175 | 2.166587 | 5.0866084 | 5 |
| CGCTG | 161430 | 2.143114 | 5.514073 | 90-94 |
| CGACG | 156455 | 2.1175108 | 5.892729 | 95-97 |
| CTGCC | 164705 | 2.1066544 | 5.596598 | 90-94 |
| GCTTT | 351885 | 2.0606687 | 6.1217313 | 1 |
| CACAT | 349195 | 2.0476296 | 6.466572 | 7 |
| CCGAC | 150915 | 1.9678597 | 5.424321 | 95-97 |
| TGCCG | 145015 | 1.9251913 | 5.5397844 | 95-97 |
| GCCAA | 212975 | 1.9156936 | 6.09992 | 1 |
| TTGGC | 207930 | 1.8678355 | 5.860738 | 5 |
| TGGCT | 206310 | 1.8532828 | 6.3949447 | 6 |
| ATACA | 441075 | 1.7841504 | 5.599916 | 4 |
| GAGTC | 191005 | 1.7492076 | 10.41032 | 9 |
| GGCTT | 188845 | 1.6963948 | 5.407702 | 1 |
| TTGAG | 272675 | 1.6896721 | 6.533209 | 9 |
| GAGAG | 173410 | 1.6804302 | 5.488952 | 7 |
| CACCA | 192295 | 1.6664447 | 5.2028804 | 7 |
| GTGTA | 258840 | 1.6039414 | 12.739067 | 1 |
| AAGAC | 256015 | 1.588541 | 6.4771485 | 5 |
| CATGA | 254590 | 1.5495272 | 6.5408998 | 9 |
| ATGGA | 239735 | 1.5144806 | 5.595535 | 4 |
| GACTT | 250450 | 1.4952149 | 8.097955 | 7 |
| TATGA | 362770 | 1.4939969 | 7.3614664 | 4 |
| GTCTT | 255100 | 1.4938874 | 7.3460765 | 1 |
| TGTGT | 244355 | 1.4852623 | 5.2116356 | 8 |
| CCATG | 168280 | 1.4847548 | 5.87065 | 9 |
| TGAGT | 239165 | 1.4820223 | 7.521905 | 8 |
| TCATA | 365380 | 1.4497349 | 7.4907956 | 2 |
| TATAC | 363005 | 1.4403114 | 5.7316155 | 5 |
| ACTCA | 241815 | 1.4179685 | 5.7273364 | 8 |
| ATGAG | 220965 | 1.3959047 | 6.617383 | 7 |
| TCCAT | 241775 | 1.3906554 | 5.6583185 | 2 |
| GTGTT | 227755 | 1.3843623 | 6.4454246 | 1 |
| AACAC | 229115 | 1.369658 | 5.6111016 | 5 |
| GATTG | 212115 | 1.3144028 | 7.3294125 | 7 |
| ACACC | 145485 | 1.2607852 | 5.673434 | 6 |
| GTTCT | 211620 | 1.239265 | 5.1303234 | 1 |
| TGGAC | 134335 | 1.2302285 | 8.897872 | 5 |
| ACCAT | 207050 | 1.2141117 | 5.221147 | 8 |
| ACATG | 198045 | 1.2053738 | 6.650111 | 8 |
| GTCCA | 136410 | 1.2035619 | 9.540075 | 1 |
| ACACT | 202295 | 1.186229 | 5.9033704 | 6 |
| ATGTG | 191070 | 1.1839943 | 5.5684304 | 7 |
| GGACT | 128570 | 1.1774334 | 8.908753 | 6 |
| ATTGA | 273855 | 1.1278179 | 5.080954 | 8 |
| AGACT | 182215 | 1.1090267 | 6.357595 | 6 |
| GTATG | 175940 | 1.090239 | 6.952461 | 3 |
| GAGTA | 170550 | 1.0774175 | 5.6477723 | 1 |
| TGTAT | 266585 | 1.0769085 | 5.945248 | 2 |
| CCATA | 179060 | 1.0499823 | 5.0971475 | 3 |
| CATAC | 174540 | 1.0234776 | 6.3401456 | 3 |
| GTATA | 198590 | 0.81785387 | 7.201834 | 1 |