![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-101_TAAGGCGA-GTAAGGAG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1584621 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
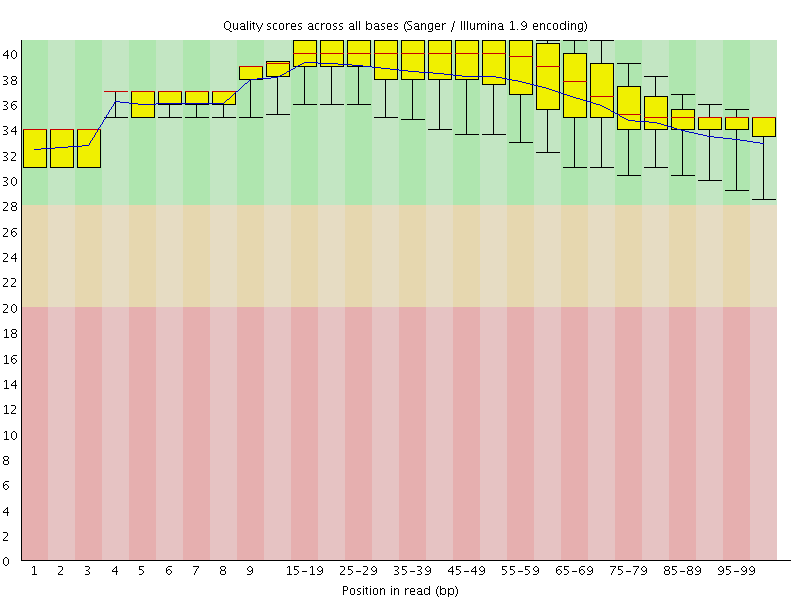
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
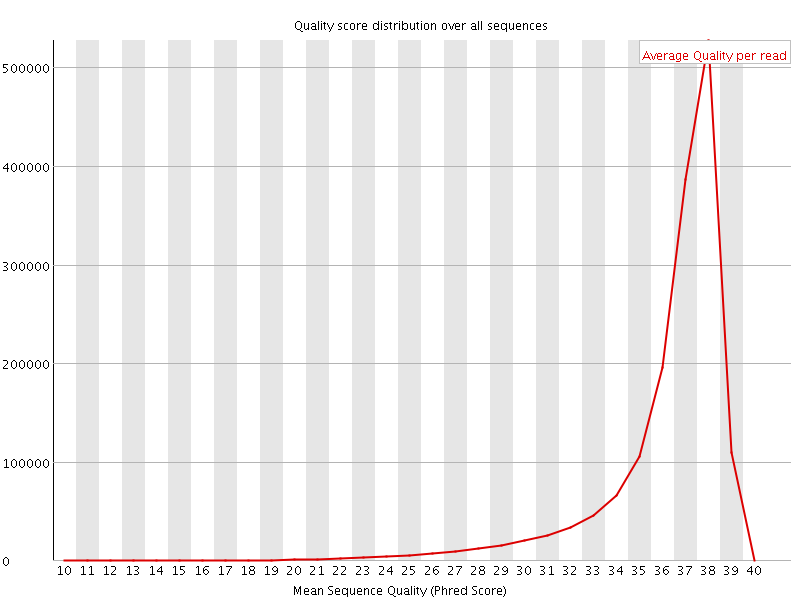
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
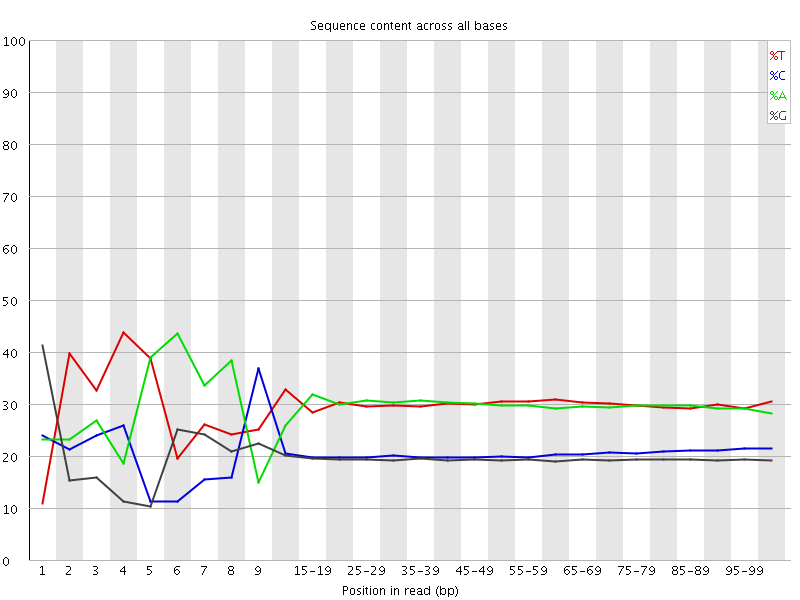
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
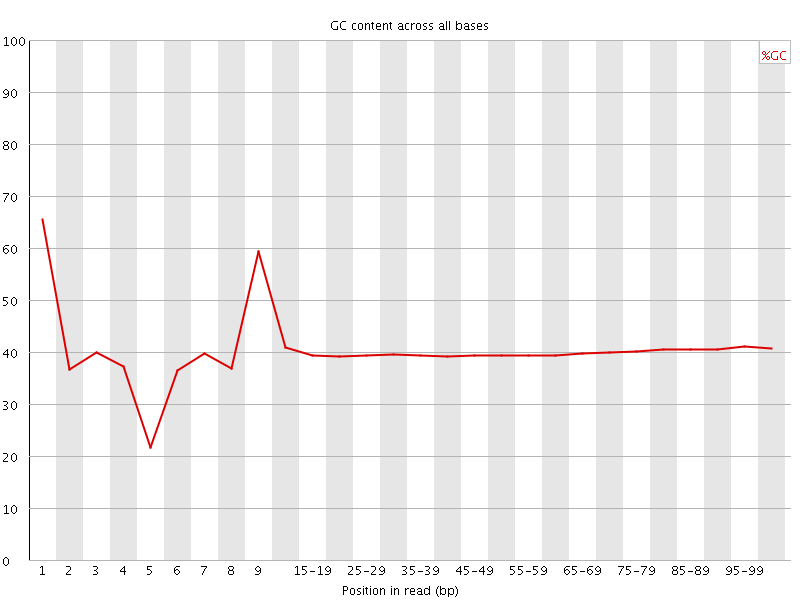
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
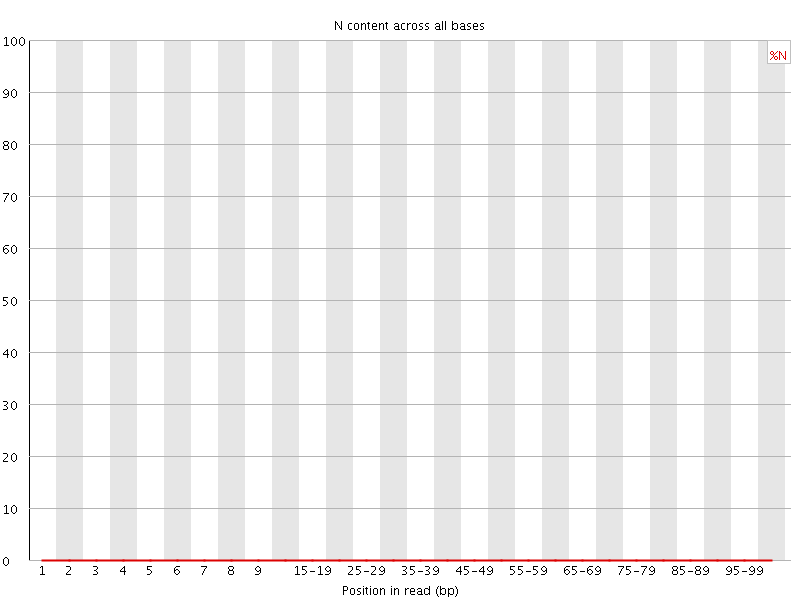
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
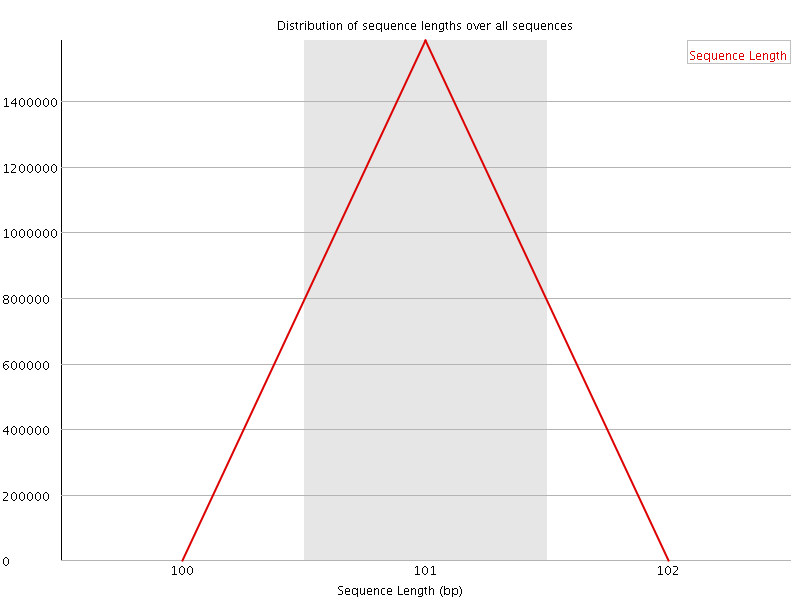
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
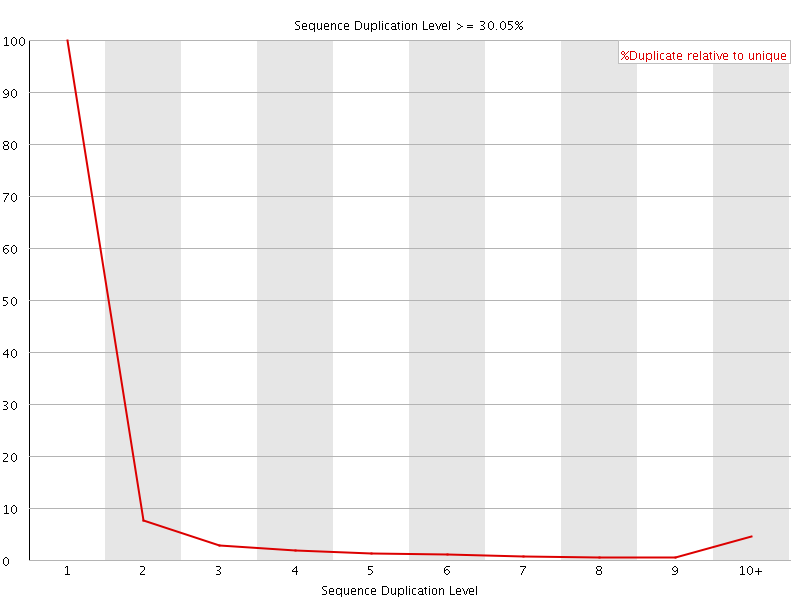
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1793 | 0.11315008446814727 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
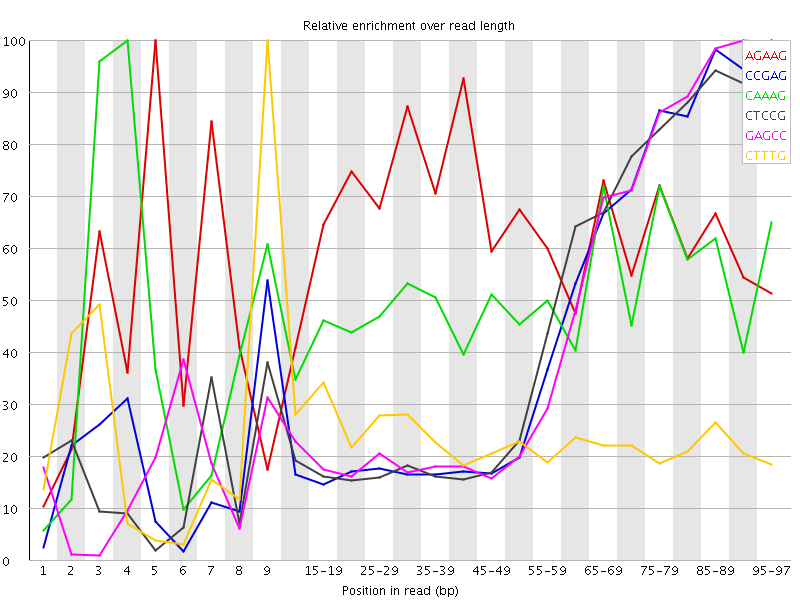
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 477940 | 3.0551763 | 4.8348784 | 5 |
| CCGAG | 199925 | 2.6900706 | 6.208619 | 95-97 |
| CAAAG | 432990 | 2.6493661 | 5.3161864 | 4 |
| CTCCG | 205840 | 2.6251686 | 5.9341903 | 95-97 |
| GAGCC | 191380 | 2.5750942 | 5.8860292 | 90-94 |
| CTTTG | 424990 | 2.5248275 | 10.645916 | 9 |
| ACACA | 419710 | 2.4581854 | 9.018699 | 6 |
| CGAGC | 180990 | 2.4352927 | 5.8367543 | 95-97 |
| TTTGG | 379505 | 2.3554251 | 5.06447 | 4 |
| GCCCA | 181565 | 2.33846 | 5.8014493 | 90-94 |
| AGCCC | 180620 | 2.326289 | 5.639084 | 90-94 |
| TACAC | 383780 | 2.225755 | 11.888527 | 5 |
| CCCAC | 174235 | 2.1480007 | 5.455097 | 90-94 |
| GCTTT | 351375 | 2.0874872 | 6.1101136 | 1 |
| CACAT | 349495 | 2.0269167 | 6.74283 | 7 |
| CCACG | 154195 | 1.985949 | 5.438 | 90-94 |
| GCCAA | 215755 | 1.9153222 | 6.108281 | 1 |
| TTGGC | 207890 | 1.8904821 | 5.6567073 | 5 |
| TGGCT | 205215 | 1.8661566 | 6.2827806 | 6 |
| GGTTG | 192425 | 1.8280969 | 5.1404366 | 7 |
| GAGTC | 192955 | 1.7720069 | 9.795579 | 9 |
| GACTC | 201105 | 1.7678013 | 6.5122643 | 7 |
| ATACA | 441155 | 1.7634753 | 5.8123627 | 4 |
| TTGAG | 276430 | 1.732636 | 6.922675 | 9 |
| GGCTT | 187550 | 1.7055169 | 5.9308496 | 1 |
| AGGCG | 120770 | 1.6976745 | 5.3548493 | 95-97 |
| GAGAG | 172580 | 1.6721257 | 5.406944 | 7 |
| GGCGA | 118610 | 1.6673111 | 5.2321672 | 95-97 |
| CACCA | 194550 | 1.6531544 | 5.4711456 | 7 |
| AGACT | 266335 | 1.6136969 | 6.257076 | 6 |
| AAGAC | 257535 | 1.5757974 | 6.3426375 | 5 |
| GTCTT | 262825 | 1.56142 | 7.2364955 | 1 |
| CATGA | 256100 | 1.551684 | 6.8886585 | 9 |
| TGTGT | 246985 | 1.5329301 | 5.449646 | 8 |
| ATGGA | 241765 | 1.5303332 | 5.450465 | 4 |
| GACTT | 253100 | 1.5185026 | 8.170961 | 7 |
| TATGA | 366010 | 1.5135597 | 7.334126 | 4 |
| CCATG | 171935 | 1.5113841 | 6.52505 | 9 |
| TGAGT | 240110 | 1.5049858 | 7.1323614 | 8 |
| TCATA | 365630 | 1.44727 | 7.575835 | 2 |
| TATAC | 365020 | 1.4448556 | 5.776601 | 5 |
| GTGTT | 231090 | 1.4342767 | 6.5699363 | 1 |
| ACTCA | 244745 | 1.419413 | 5.7418094 | 8 |
| GTGTA | 225995 | 1.4165144 | 13.129904 | 1 |
| TCCAT | 244560 | 1.4044621 | 5.173994 | 2 |
| ATGAG | 221455 | 1.4017739 | 6.113359 | 7 |
| AACAC | 227025 | 1.3296549 | 5.4208736 | 5 |
| GATTG | 211925 | 1.3283249 | 7.3329306 | 7 |
| GTATG | 204010 | 1.2787145 | 6.9135585 | 3 |
| ACACC | 146535 | 1.2451555 | 6.0767627 | 6 |
| GTTCT | 209525 | 1.2447692 | 5.31214 | 1 |
| TGGAC | 135135 | 1.2410154 | 8.753685 | 5 |
| ACCAT | 210445 | 1.2204881 | 5.4493766 | 8 |
| GTCCA | 137360 | 1.2074547 | 8.601761 | 1 |
| ATGTG | 191265 | 1.1988301 | 5.40929 | 7 |
| ACATG | 197775 | 1.1982989 | 6.982662 | 8 |
| GGACT | 129315 | 1.1875674 | 8.718065 | 6 |
| ACACT | 201965 | 1.171308 | 5.9245806 | 6 |
| ATTGA | 273970 | 1.132947 | 5.2569923 | 8 |
| TGTAT | 267430 | 1.0950811 | 6.0818934 | 2 |
| GAGTA | 171560 | 1.0859468 | 5.3560367 | 1 |
| TGTAG | 164990 | 1.0341411 | 5.2641187 | 2 |
| CATAC | 174360 | 1.011211 | 6.433526 | 3 |
| GTATA | 198885 | 0.82244825 | 7.242857 | 1 |