![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL5_CAGATC_L008_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9436412 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
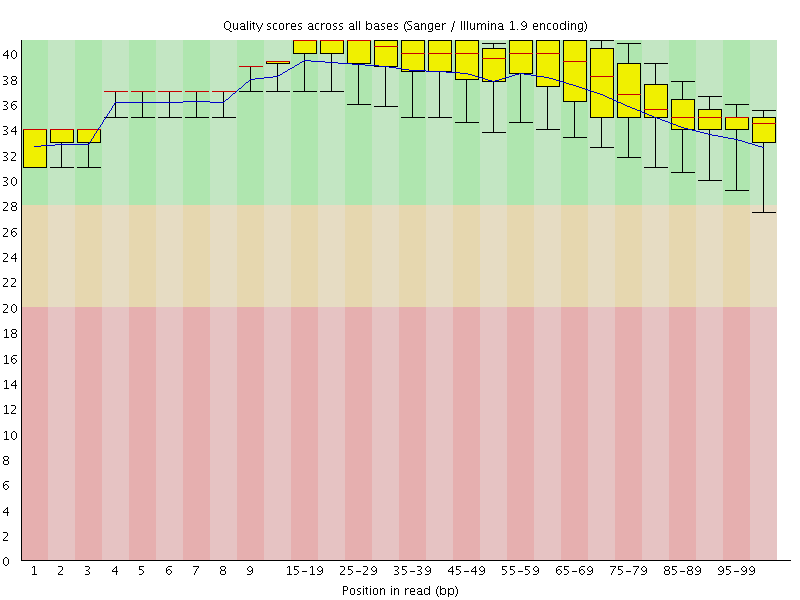
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
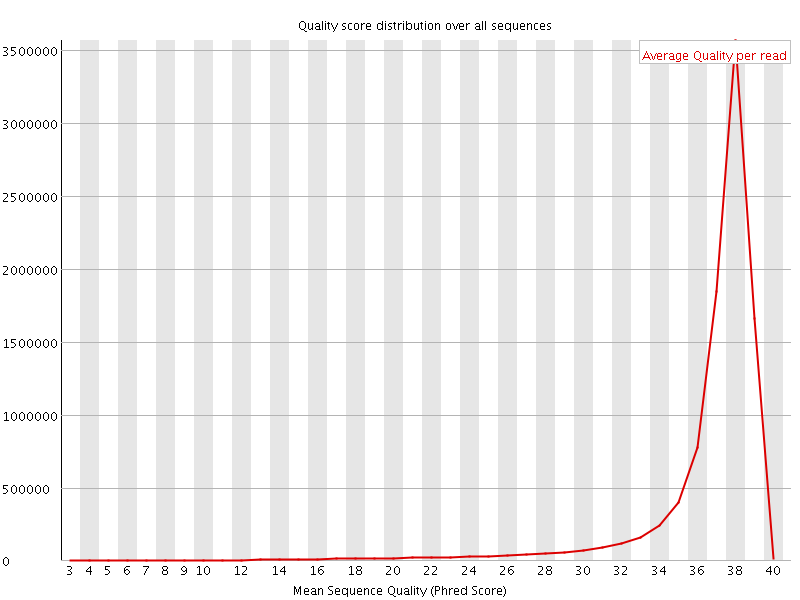
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
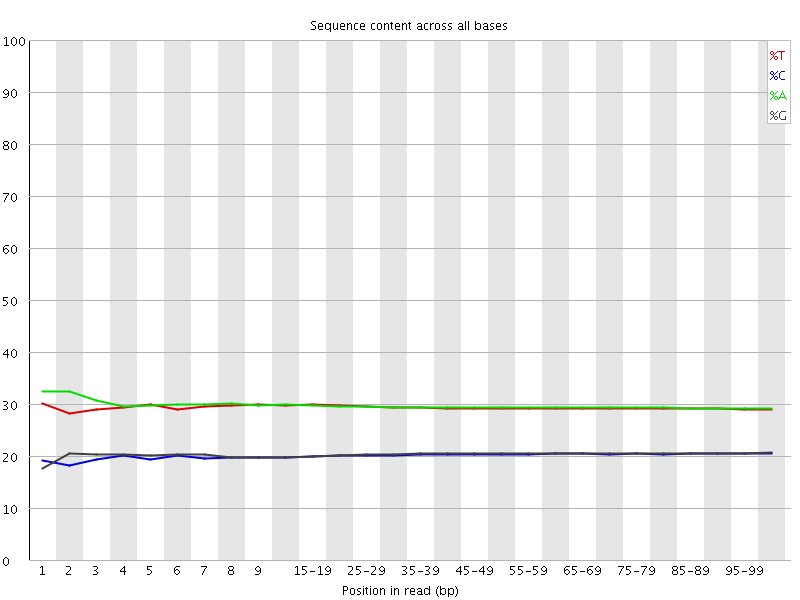
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
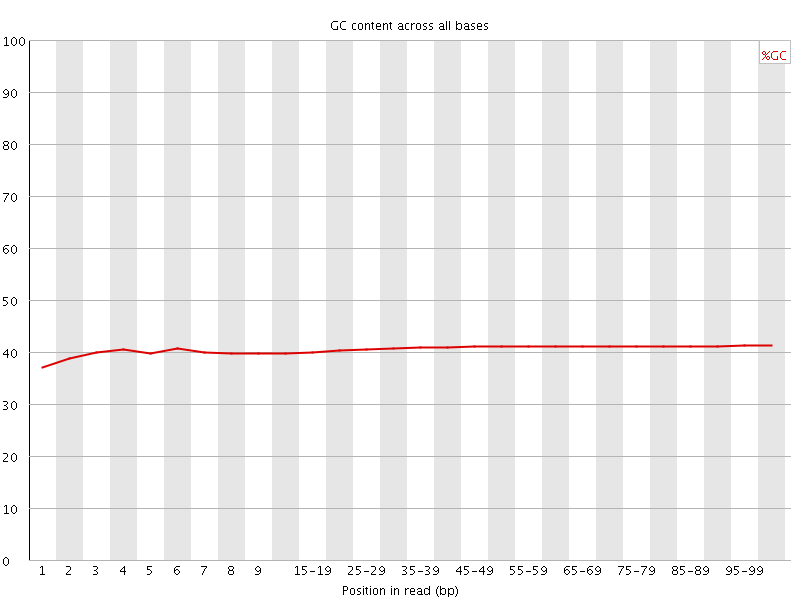
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
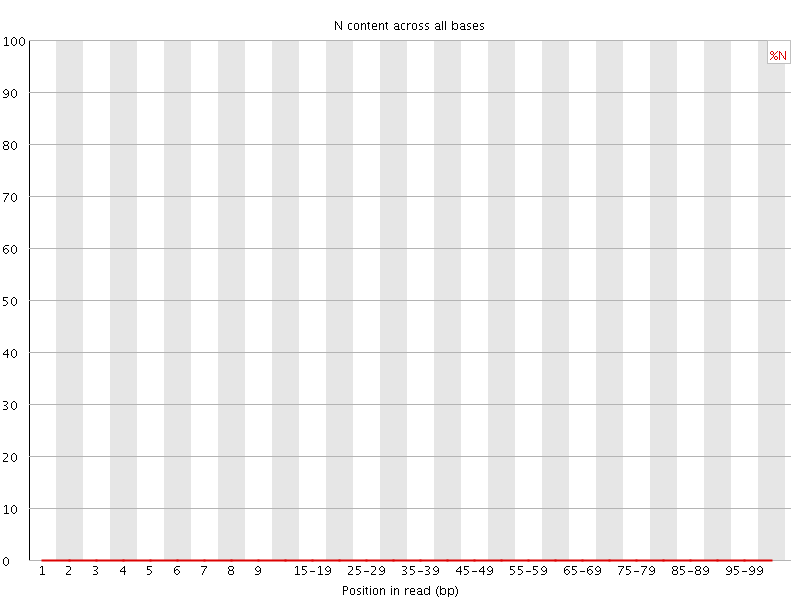
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
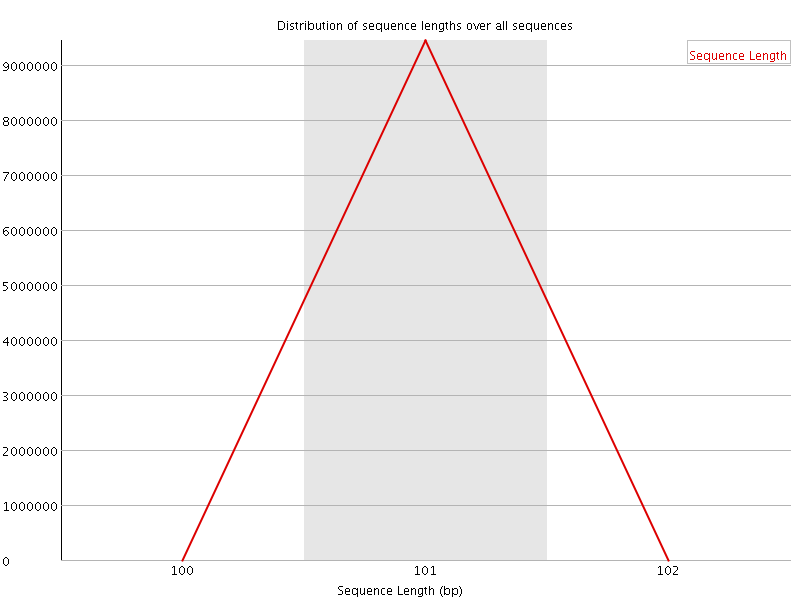
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
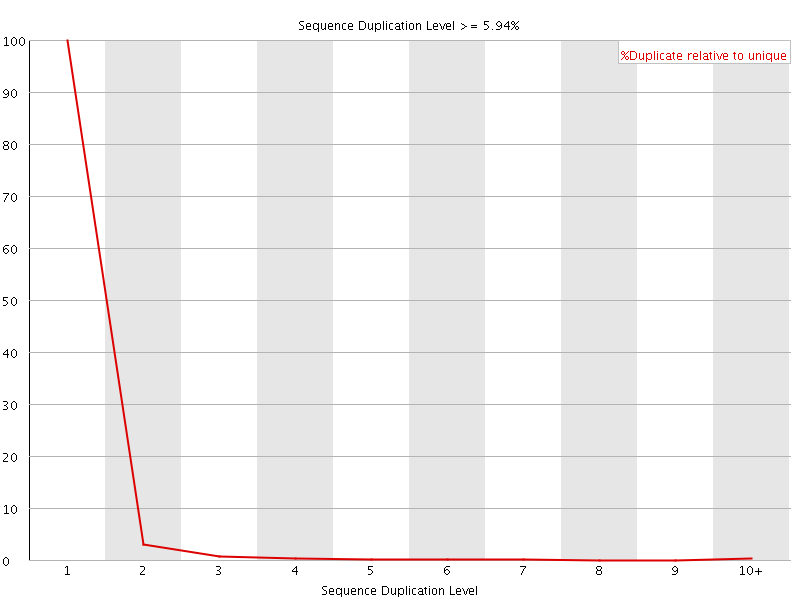
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
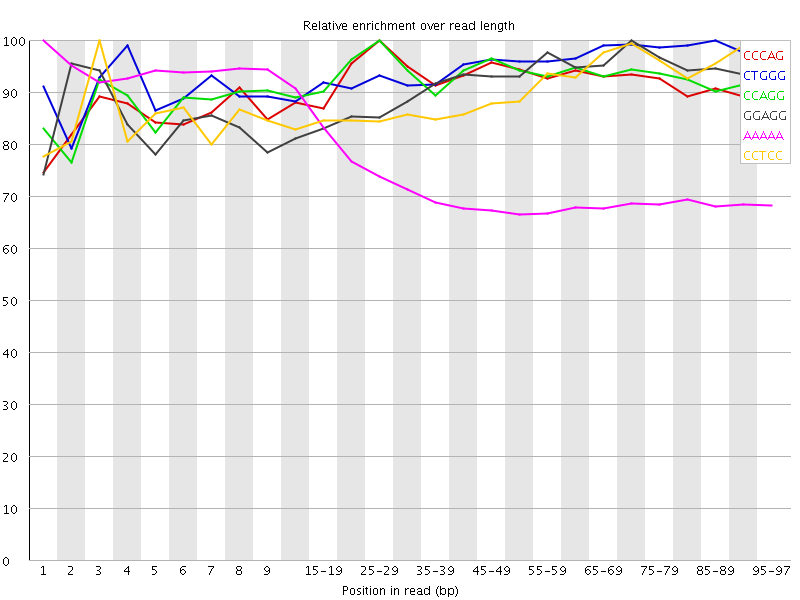
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCCAG | 1695945 | 3.5997813 | 3.9122376 | 25-29 |
| CTGGG | 1660065 | 3.4995422 | 3.6845202 | 85-89 |
| CCAGG | 1514365 | 3.1924126 | 3.439897 | 25-29 |
| GGAGG | 1522300 | 3.165461 | 3.4662902 | 70-74 |
| AAAAA | 6530365 | 3.114264 | 4.239692 | 1 |
| CCTCC | 1447015 | 3.1137724 | 3.4517603 | 3 |
| CCTGG | 1461340 | 3.1017964 | 3.2829425 | 70-74 |
| GGGGG | 1027610 | 3.0877683 | 4.0836396 | 95-97 |
| TTTTT | 6199055 | 3.059224 | 3.6462307 | 10-14 |