![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL5_CAGATC_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9340932 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
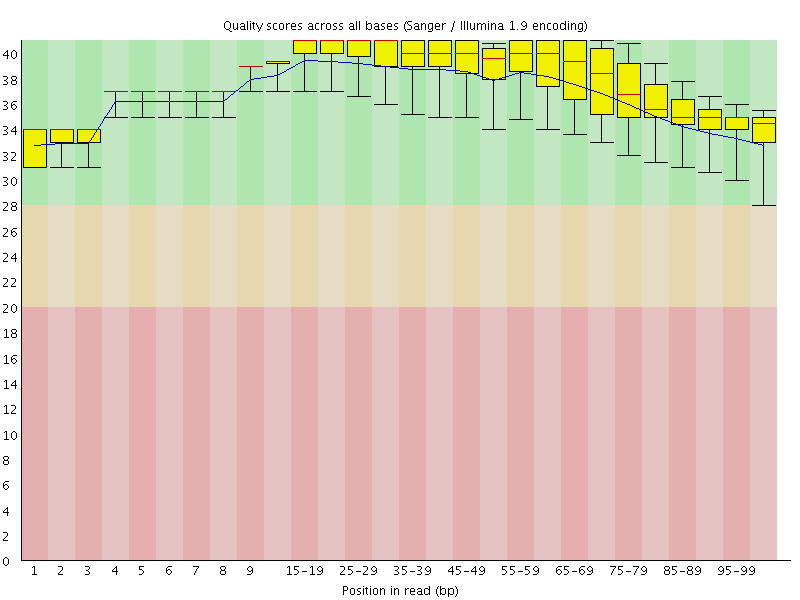
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
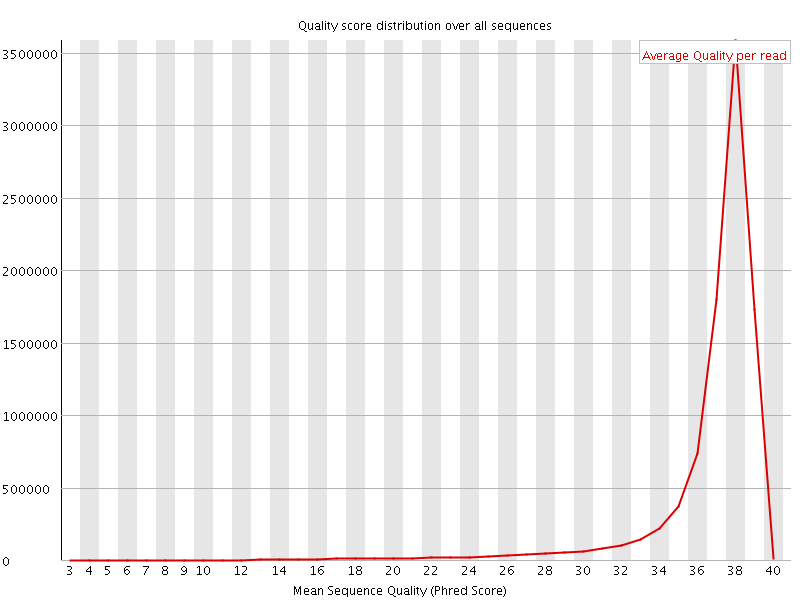
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
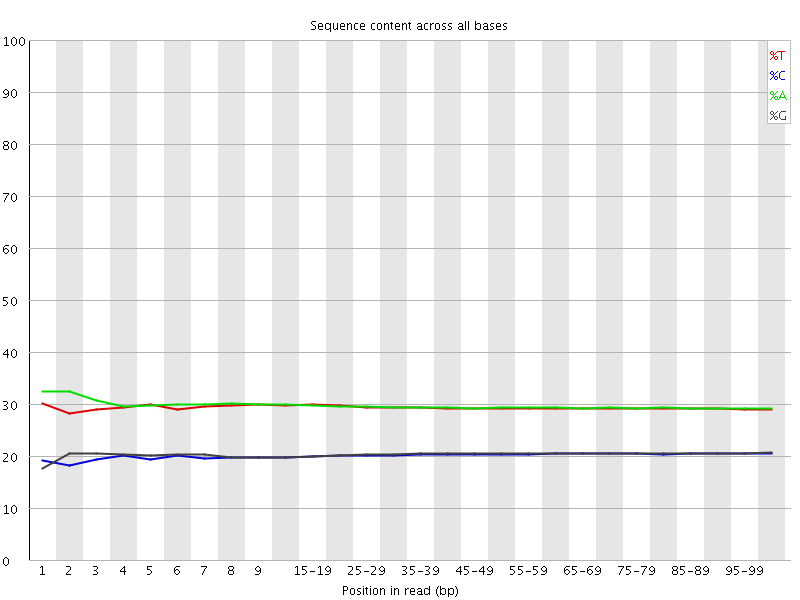
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
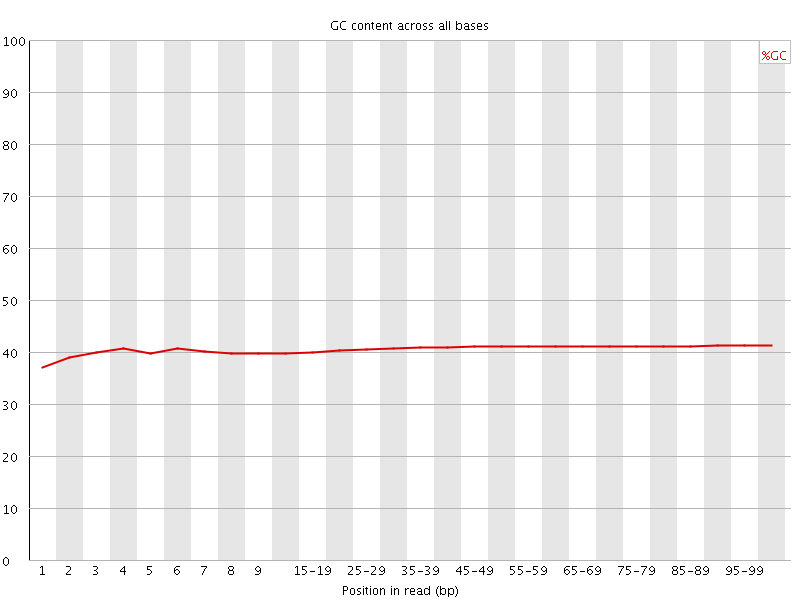
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
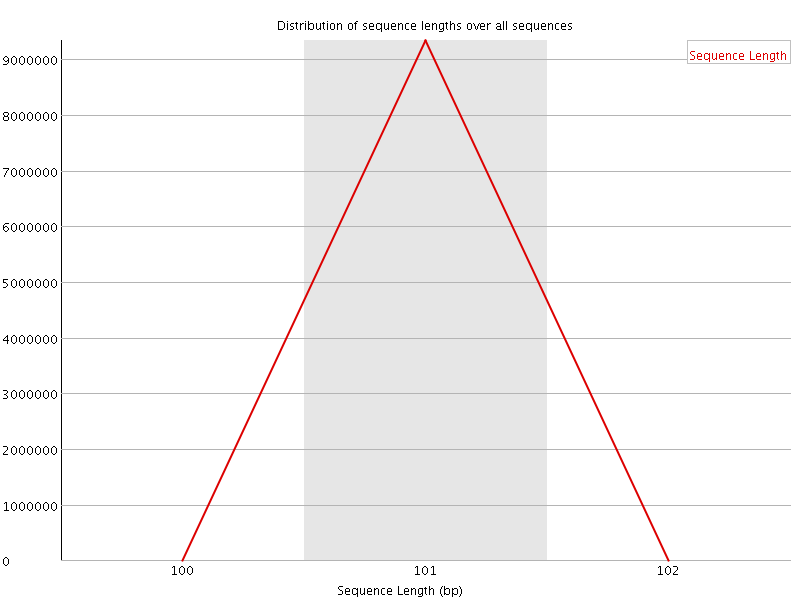
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
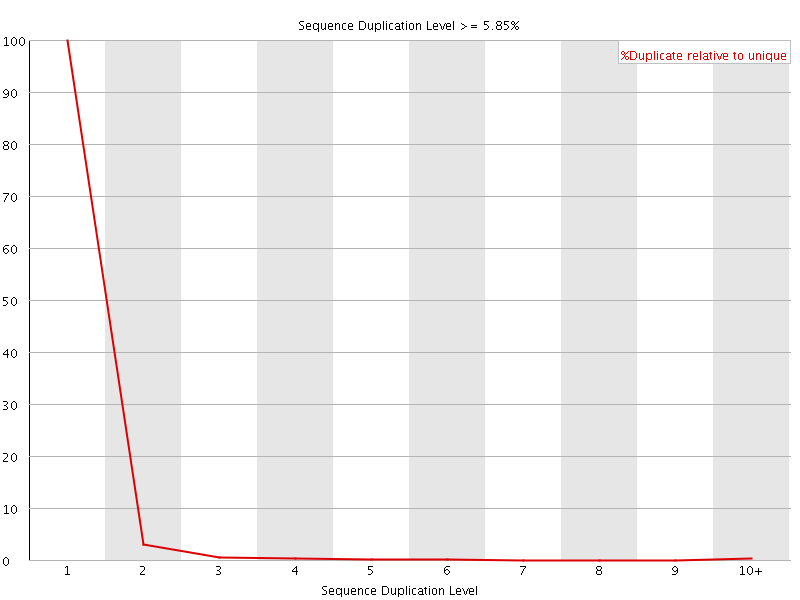
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
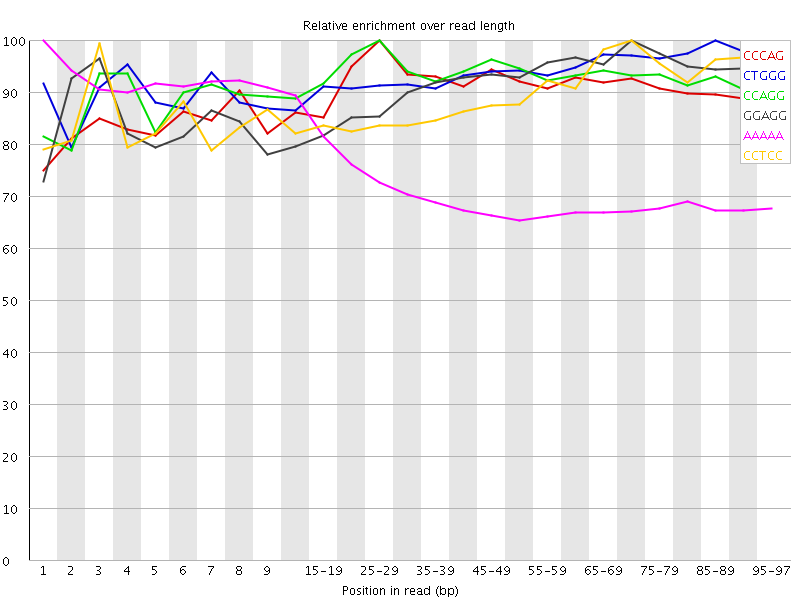
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCCAG | 1688390 | 3.6121569 | 3.974485 | 25-29 |
| CTGGG | 1643835 | 3.495128 | 3.7273588 | 85-89 |
| CCAGG | 1508815 | 3.2071579 | 3.445896 | 25-29 |
| GGAGG | 1507455 | 3.1630743 | 3.4600632 | 70-74 |
| AAAAA | 6484175 | 3.1300828 | 4.320057 | 1 |
| CCTCC | 1432940 | 3.106431 | 3.473019 | 70-74 |
| CCTGG | 1448345 | 3.0994632 | 3.3106866 | 65-69 |
| GGGGG | 1019430 | 3.0886126 | 4.1241837 | 95-97 |
| TTTTT | 6149585 | 3.0704212 | 3.6685016 | 10-14 |