![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L008_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11041035 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
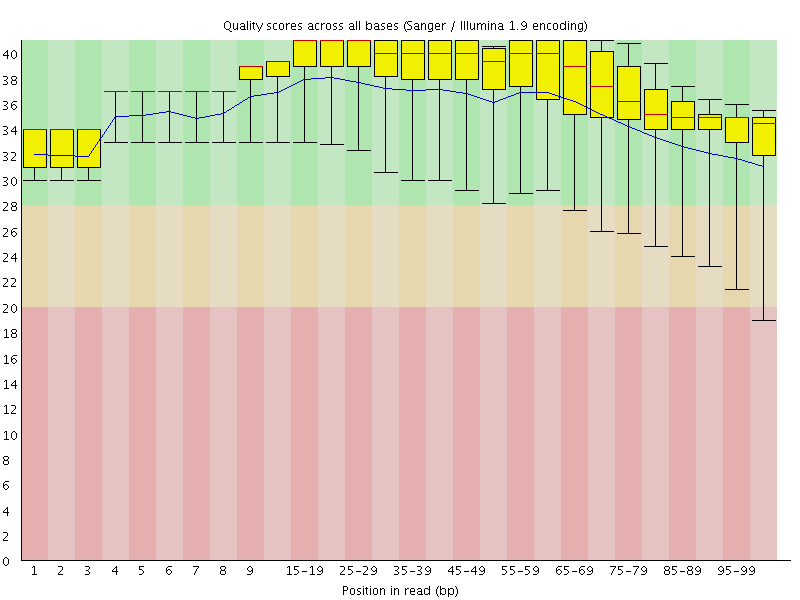
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
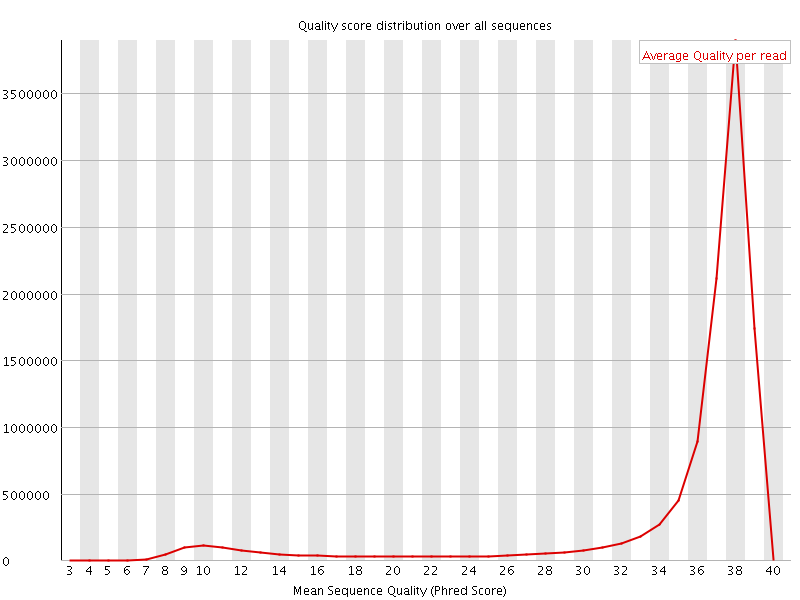
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
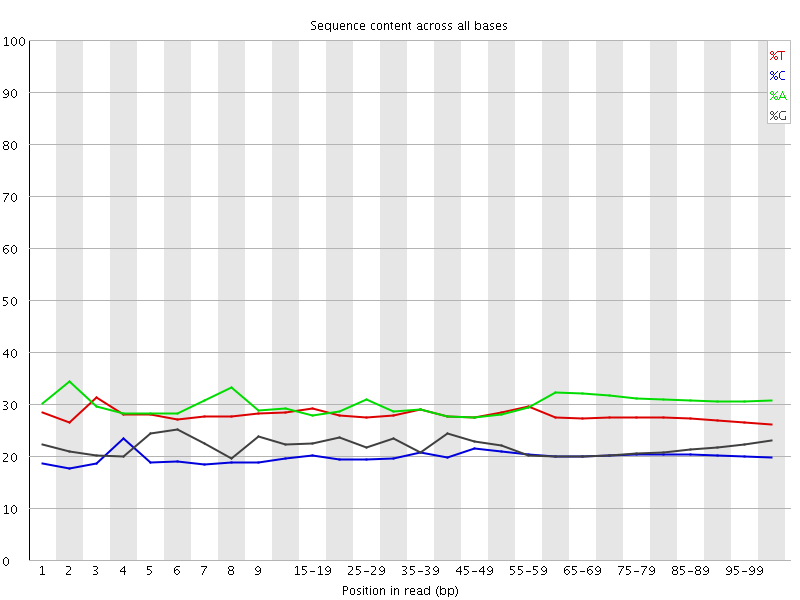
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
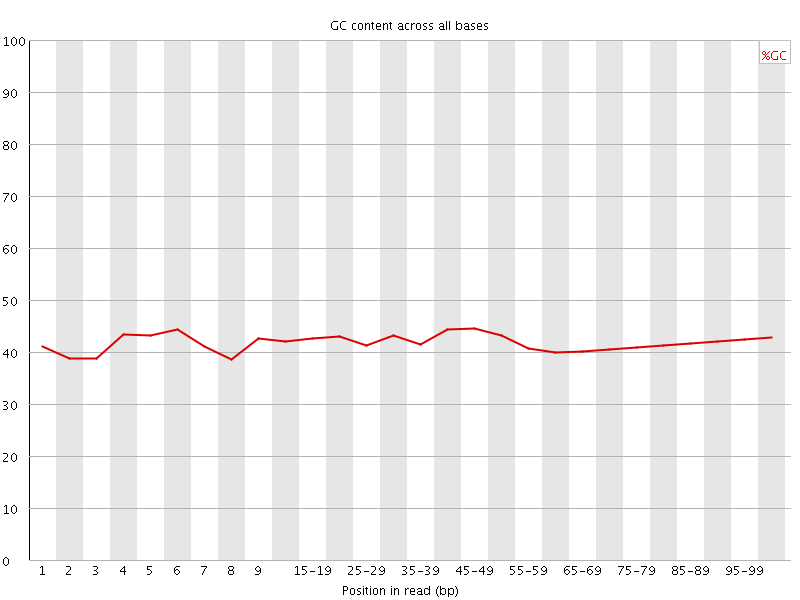
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
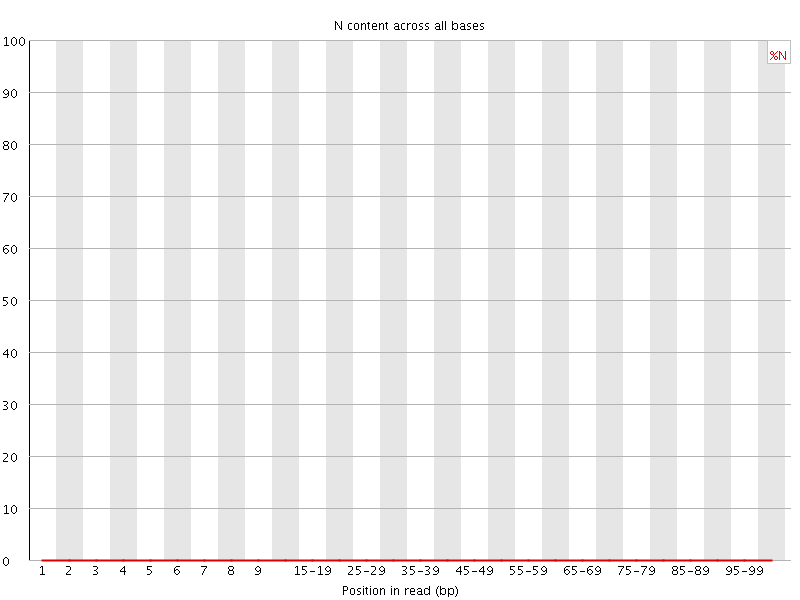
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
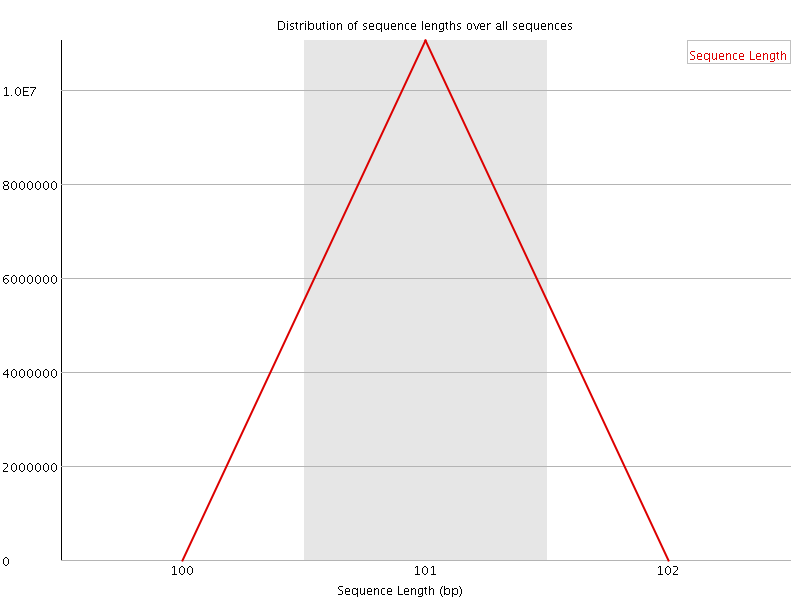
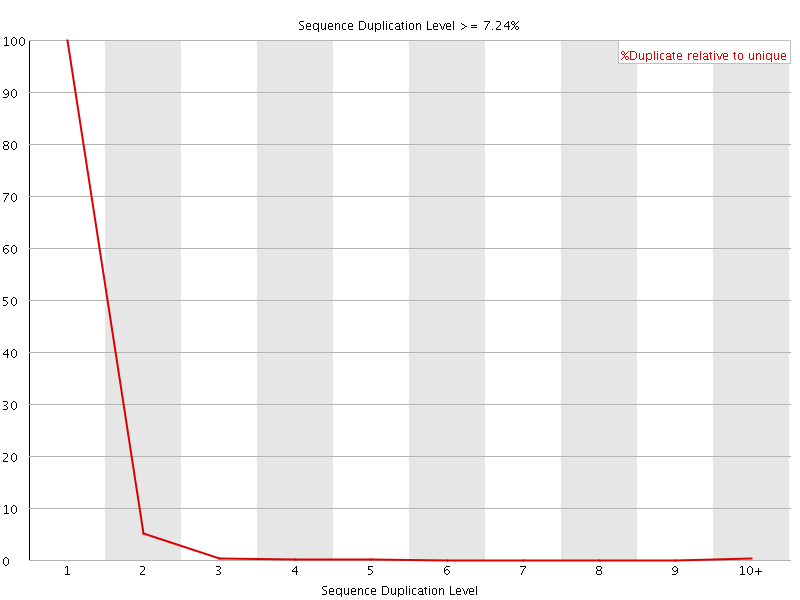
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCTCGGTGGTCGCCG | 14088 | 0.1275967334584122 | Illumina Single End PCR Primer 1 (100% over 50bp) |
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGGGTAGATCTCGGGGGGCGCCG | 11362 | 0.10290701913362289 | Illumina Single End PCR Primer 1 (97% over 41bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
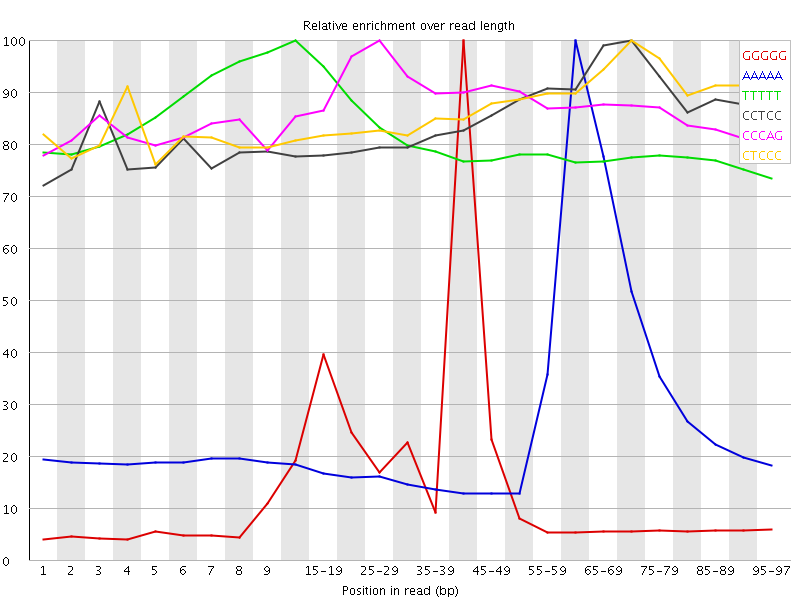
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 4473355 | 8.312492 | 49.855194 | 40-44 |
| AAAAA | 14360830 | 5.530773 | 19.476723 | 60-64 |
| TTTTT | 6860210 | 3.7638187 | 4.634643 | 10-14 |
| CCTCC | 1776945 | 3.5837615 | 4.188521 | 70-74 |
| CCCAG | 2047125 | 3.545605 | 4.0361514 | 25-29 |
| CTCCC | 1587785 | 3.2022617 | 3.659502 | 70-74 |
| CTGGG | 2012450 | 3.1786582 | 3.4054868 | 70-74 |
| GGGAG | 2249945 | 3.0519083 | 23.983824 | 5 |
| CCTGG | 1760445 | 3.0166388 | 3.2783172 | 70-74 |
| GGAGG | 2180115 | 2.9571884 | 5.457682 | 6 |
| GGAAG | 2672930 | 2.6466033 | 27.317019 | 5 |
| GGGGC | 1302495 | 2.6257665 | 10.242613 | 40-44 |
| GGGGA | 1928855 | 2.61637 | 11.258057 | 20-24 |
| AGGGG | 1852115 | 2.5122774 | 9.167907 | 25-29 |
| GAGGG | 1737410 | 2.356687 | 15.6213875 | 9 |
| GGGAA | 2141135 | 2.1200461 | 11.316214 | 20-24 |
| AAGAG | 2917995 | 2.1090531 | 19.684778 | 7 |
| GAAGA | 2754245 | 1.9906988 | 20.068624 | 6 |
| GGAGA | 2002850 | 1.9831232 | 18.309193 | 6 |
| AGAGC | 1833120 | 1.9691298 | 34.088276 | 8 |
| AGAGG | 1921410 | 1.9024855 | 11.266047 | 8 |
| GGGCC | 852590 | 1.864672 | 6.2340603 | 45-49 |
| AGGGA | 1827135 | 1.809139 | 5.0117297 | 20-24 |
| GGAAA | 2459225 | 1.7774656 | 6.0190883 | 20-24 |
| GAGAG | 1753080 | 1.7358134 | 17.468744 | 7 |
| AGATC | 1976360 | 1.6633673 | 6.5515356 | 95-97 |
| CGGGG | 818755 | 1.6505704 | 9.218188 | 35-39 |
| GATCG | 1394120 | 1.6073875 | 41.966675 | 1 |
| ATCGG | 1305070 | 1.504715 | 41.883865 | 2 |
| GGGCG | 706990 | 1.4252576 | 6.0121393 | 40-44 |
| GGCGG | 698150 | 1.4074366 | 7.224037 | 10-14 |
| GAGCG | 925830 | 1.3624263 | 45.387794 | 9 |
| TCGGA | 1108610 | 1.2782011 | 28.807758 | 3 |
| CGGAA | 1122095 | 1.2053497 | 28.00831 | 4 |
| GCGTC | 674215 | 1.1553121 | 7.465699 | 10-14 |
| TAGGG | 1085340 | 1.1534642 | 6.473061 | 15-19 |
| GCGGG | 568800 | 1.1466732 | 6.879048 | 10-14 |
| GTAGG | 1022655 | 1.0868447 | 6.4594574 | 15-19 |
| CGTCG | 556485 | 0.95357394 | 7.3493333 | 10-14 |
| TCGGG | 592840 | 0.93638885 | 20.291002 | 3 |
| GCCGG | 419380 | 0.9172123 | 6.942408 | 45-49 |
| CTCGG | 515330 | 0.883052 | 6.249949 | 35-39 |
| AGCGT | 744865 | 0.85881174 | 5.0406475 | 10-14 |
| GGCGC | 391800 | 0.85689306 | 5.222759 | 40-44 |
| GCGCC | 354950 | 0.8421928 | 5.611309 | 45-49 |
| CGTGT | 677190 | 0.83804625 | 5.730671 | 15-19 |
| CGGGA | 471495 | 0.69383925 | 22.094296 | 4 |
| CGCCG | 238090 | 0.5649181 | 6.823974 | 45-49 |