![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11041035 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
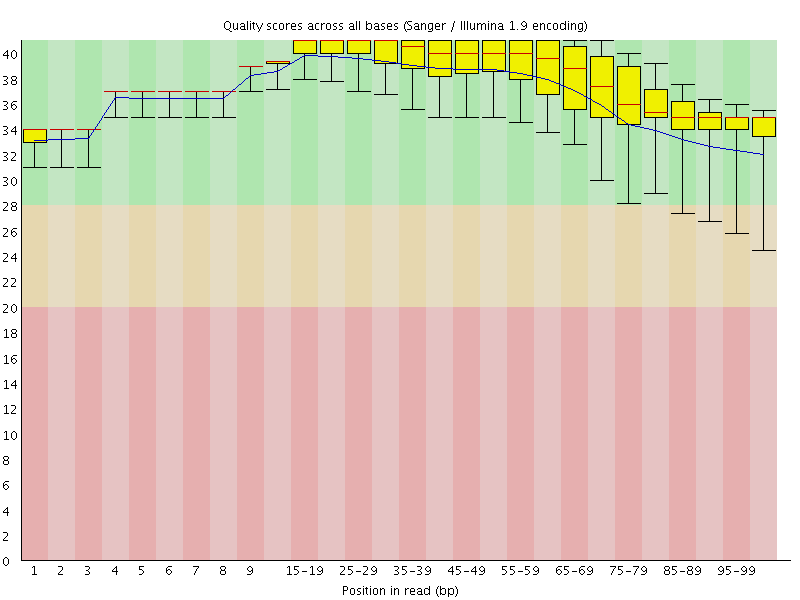
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
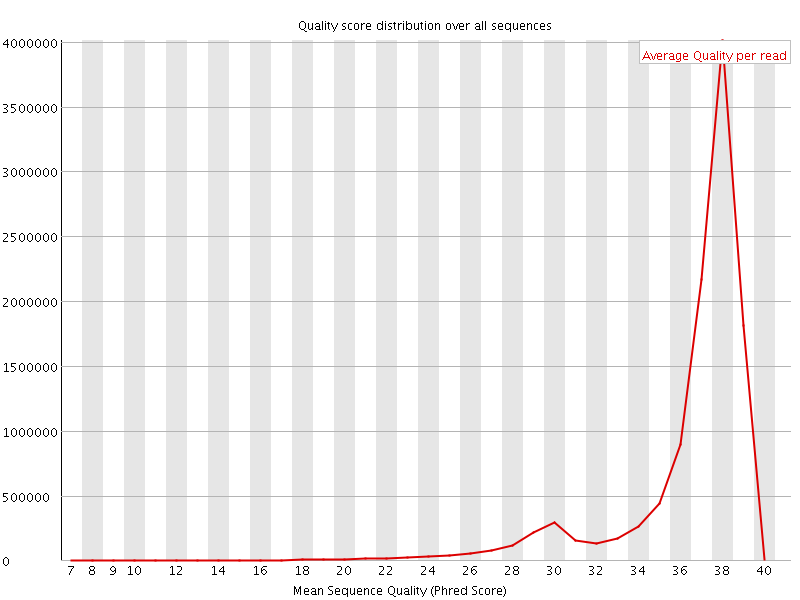
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
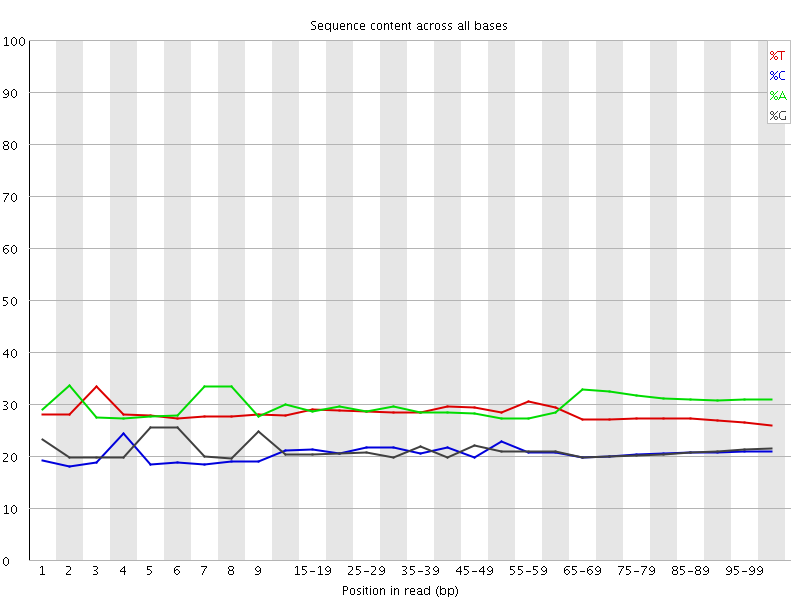
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
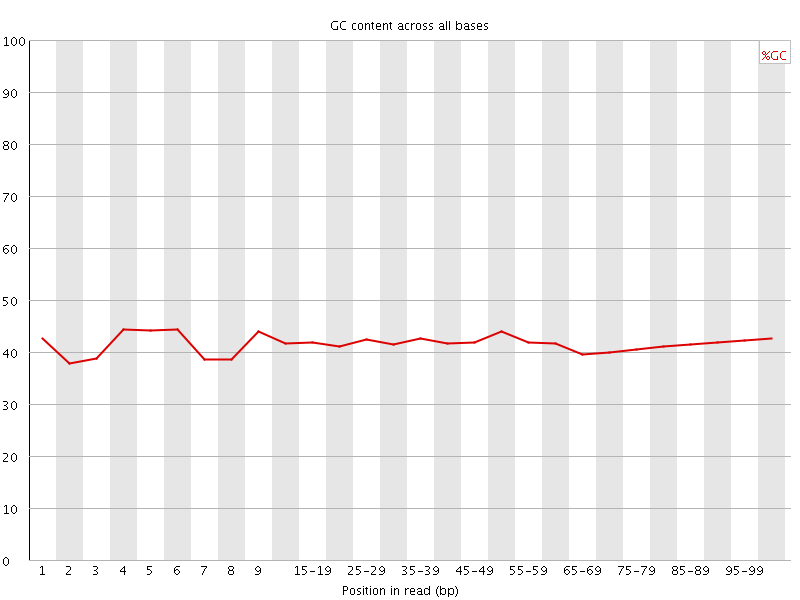
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
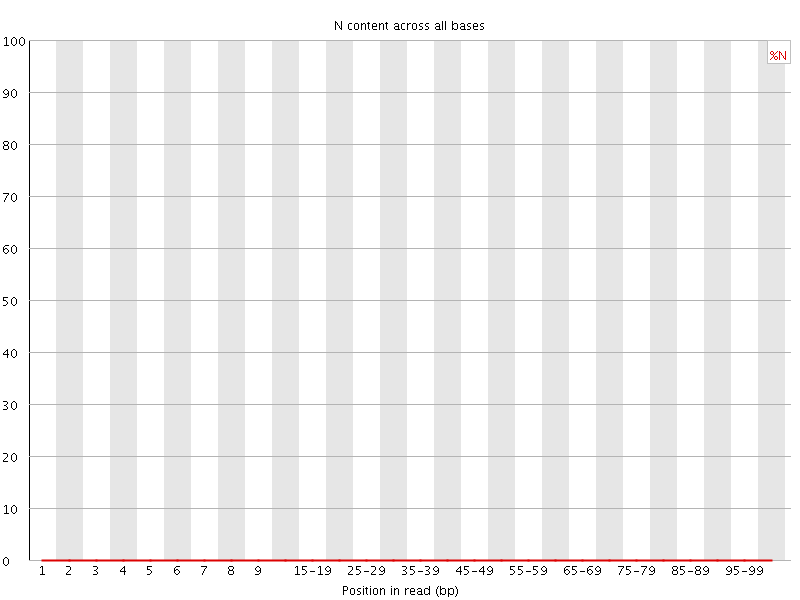
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
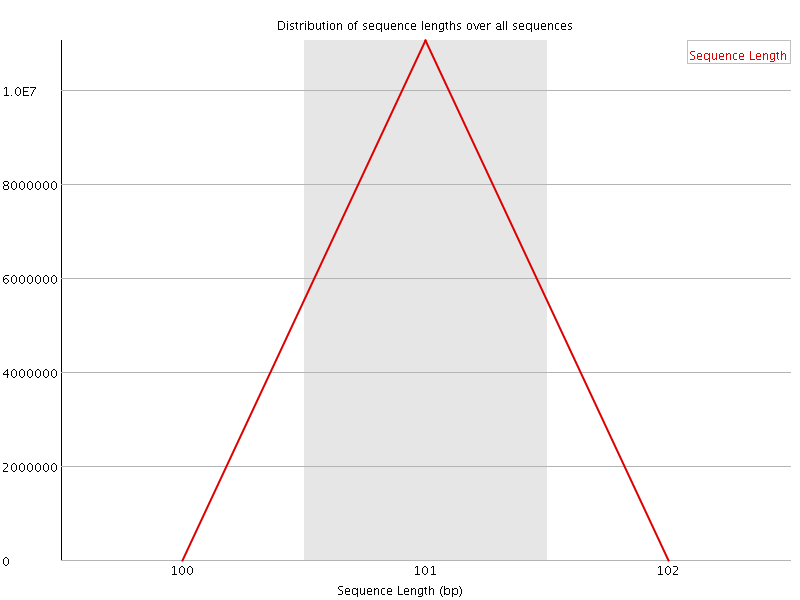
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
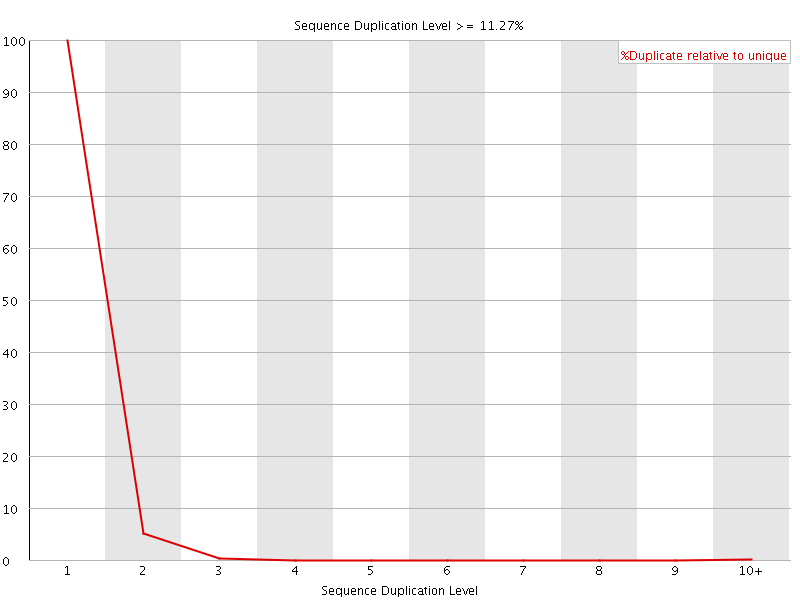
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGC | 552728 | 5.006124878691173 | TruSeq Adapter, Index 5 (100% over 50bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
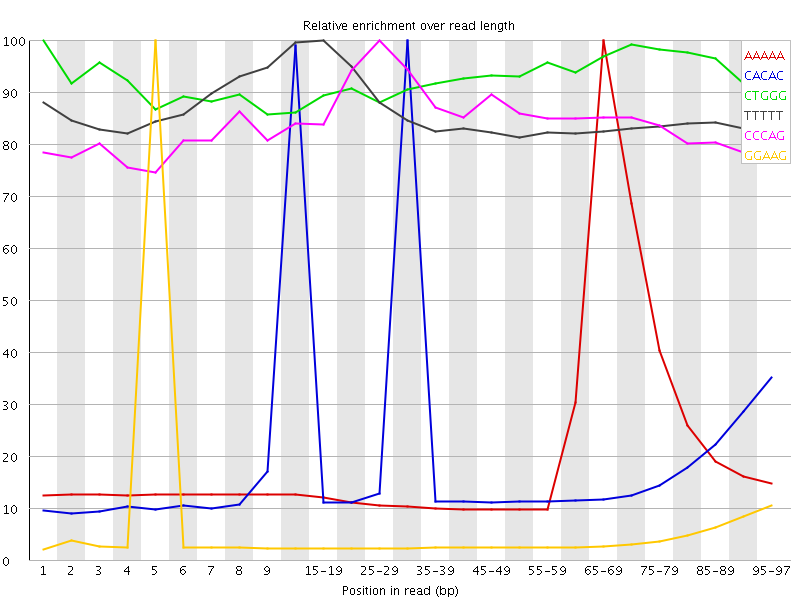
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 13876020 | 5.439093 | 23.975899 | 65-69 |
| CACAC | 3090240 | 3.5596724 | 15.284698 | 30-34 |
| CTGGG | 1995330 | 3.4500887 | 3.7105205 | 1 |
| TTTTT | 6642160 | 3.387125 | 3.9356759 | 15-19 |
| CCCAG | 2054990 | 3.3841074 | 3.957748 | 25-29 |
| GGAAG | 2730240 | 3.126887 | 69.11971 | 5 |
| CCTCC | 1785775 | 3.1056228 | 3.6086152 | 70-74 |
| CCTGG | 1765340 | 3.058295 | 3.3031144 | 65-69 |
| GGAGG | 1863195 | 3.0506072 | 3.506959 | 4 |
| CCAGG | 1830205 | 3.0081441 | 3.4754035 | 25-29 |
| CTCCA | 2192910 | 2.6625063 | 16.135422 | 20-24 |
| CAGTG | 2181365 | 2.6383188 | 15.994941 | 35-39 |
| TCCAG | 2149285 | 2.6045244 | 15.812702 | 25-29 |
| CTGCT | 1981735 | 2.5312326 | 16.859957 | 55-59 |
| CACAG | 2174495 | 2.500004 | 15.499112 | 30-34 |
| TCTGC | 1898690 | 2.4251604 | 16.743626 | 55-59 |
| AGAGC | 2105760 | 2.416327 | 68.3071 | 8 |
| TTCTG | 2484275 | 2.3349957 | 12.657498 | 55-59 |
| GAGCA | 2019455 | 2.3172932 | 68.17861 | 9 |
| GCACA | 1999720 | 2.2990663 | 14.685558 | 10-14 |
| AGCAC | 1984435 | 2.2814932 | 14.575521 | 10-14 |
| GAAGA | 2837885 | 2.2734716 | 48.361984 | 6 |
| AAGAG | 2776845 | 2.2245717 | 48.282364 | 7 |
| CTTCT | 2345640 | 2.2089367 | 12.8106575 | 50-54 |
| ACACA | 2694875 | 2.1672266 | 11.026716 | 30-34 |
| GTCTG | 1689175 | 2.153405 | 16.17256 | 15-19 |
| ACTCC | 1765740 | 2.143861 | 15.736228 | 20-24 |
| CGTCT | 1668700 | 2.1313987 | 15.322741 | 15-19 |
| TCTTC | 2156605 | 2.0309186 | 12.722144 | 50-54 |
| CCAGT | 1660310 | 2.0119796 | 15.380411 | 25-29 |
| TGCTT | 2086085 | 1.960733 | 12.696609 | 55-59 |
| GATCG | 1617810 | 1.9567099 | 71.599205 | 1 |
| TCTGA | 2156275 | 1.9228195 | 11.906881 | 15-19 |
| CTGAA | 2240410 | 1.8954389 | 11.298687 | 15-19 |
| GCTTG | 1485680 | 1.8939837 | 16.319477 | 55-59 |
| GAAAA | 3377805 | 1.8928349 | 8.109623 | 60-64 |
| ATGCC | 1560570 | 1.8911138 | 15.444479 | 45-49 |
| ATCGG | 1497890 | 1.8116688 | 71.51031 | 2 |
| AGTGA | 2113105 | 1.7843002 | 11.084857 | 35-39 |
| CAGTC | 1464120 | 1.7742345 | 15.18589 | 25-29 |
| TCACA | 2080330 | 1.7633967 | 11.035793 | 30-34 |
| TCGGA | 1450575 | 1.7544426 | 71.46219 | 3 |
| ATCTC | 1900830 | 1.6982949 | 11.302486 | 40-44 |
| CTTGA | 1901010 | 1.6951915 | 11.562041 | 60-64 |
| TGAAA | 2830890 | 1.6720654 | 8.21467 | 60-64 |
| CGGAA | 1446215 | 1.6595093 | 67.769104 | 4 |
| GTCTT | 1748020 | 1.6429822 | 12.152958 | 50-54 |
| GTCAC | 1353410 | 1.6400753 | 15.258293 | 25-29 |
| AGATC | 1818985 | 1.5389036 | 6.46109 | 95-97 |
| GTGAT | 1718850 | 1.5298078 | 11.333573 | 35-39 |
| AACTC | 1803645 | 1.5288639 | 10.894396 | 20-24 |
| TGAAC | 1800825 | 1.5235398 | 10.738726 | 20-24 |
| ACAGT | 1775385 | 1.502017 | 10.959023 | 30-34 |
| TTGAA | 2402865 | 1.4959314 | 8.444279 | 60-64 |
| GAACT | 1738400 | 1.4707268 | 10.797476 | 20-24 |
| AGTCA | 1692740 | 1.4320974 | 10.978812 | 25-29 |
| CCGTC | 815635 | 1.4157361 | 20.354841 | 50-54 |
| TGATC | 1538465 | 1.3718985 | 11.194419 | 35-39 |
| CACGT | 1128545 | 1.3675817 | 14.732805 | 10-14 |
| TGCCG | 788860 | 1.36663 | 21.08041 | 45-49 |
| GATCT | 1514225 | 1.3502829 | 11.164403 | 35-39 |
| ACACG | 1136975 | 1.3071735 | 13.970601 | 10-14 |
| GCCGT | 751825 | 1.3024701 | 20.735435 | 45-49 |
| TCTCG | 924865 | 1.1813124 | 15.162213 | 40-44 |
| ACGTC | 972475 | 1.1784545 | 14.328633 | 15-19 |
| TATGC | 1313540 | 1.1713256 | 11.091868 | 45-49 |
| GTATG | 1301800 | 1.1586257 | 10.921036 | 45-49 |
| CTCGT | 794910 | 1.0153235 | 15.0352335 | 40-44 |
| CGTAT | 769475 | 0.6861655 | 10.709678 | 40-44 |
| TCGTA | 730705 | 0.65159315 | 10.769096 | 40-44 |