![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10869007 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
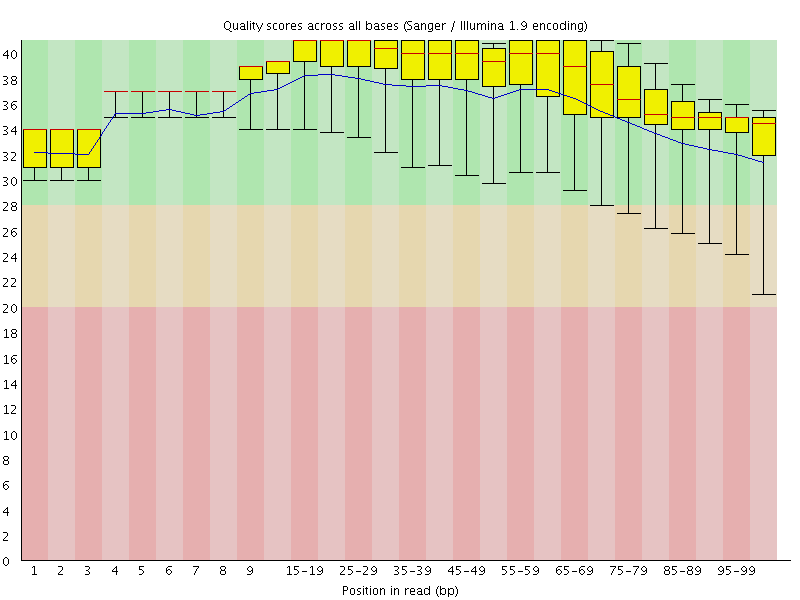
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
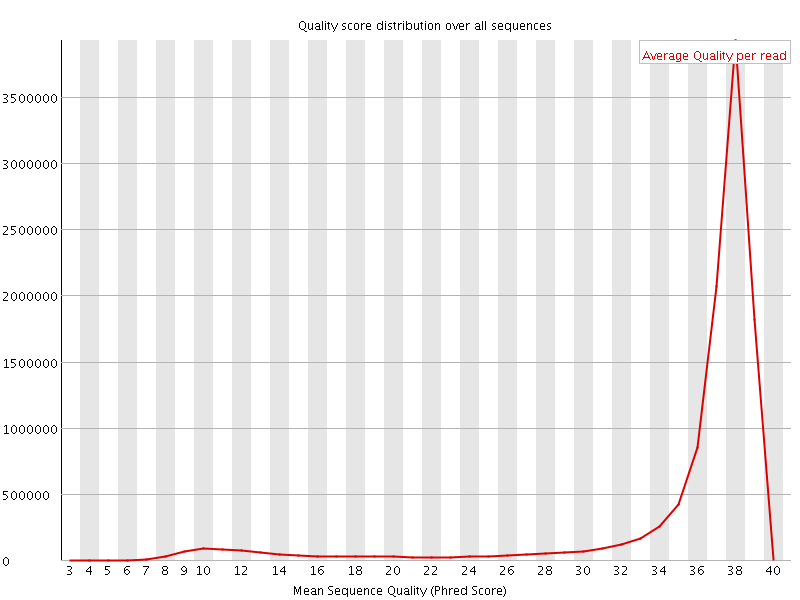
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
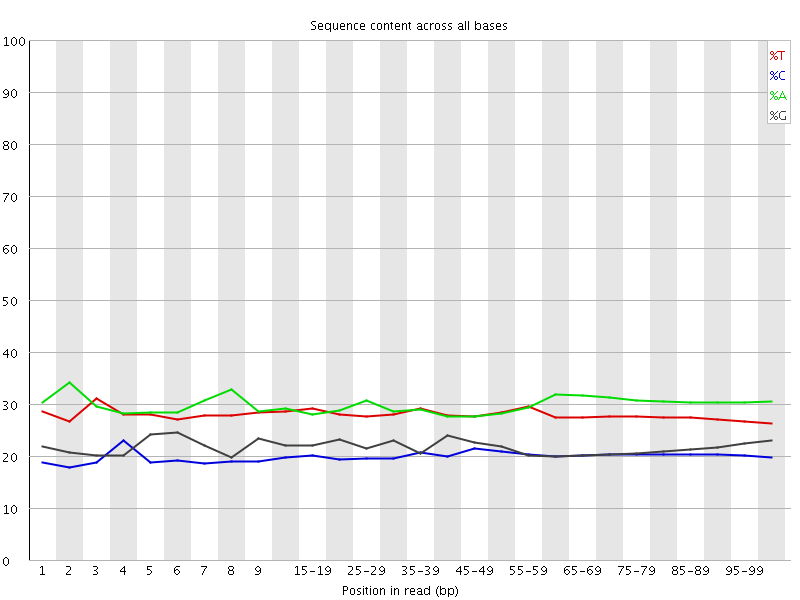
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
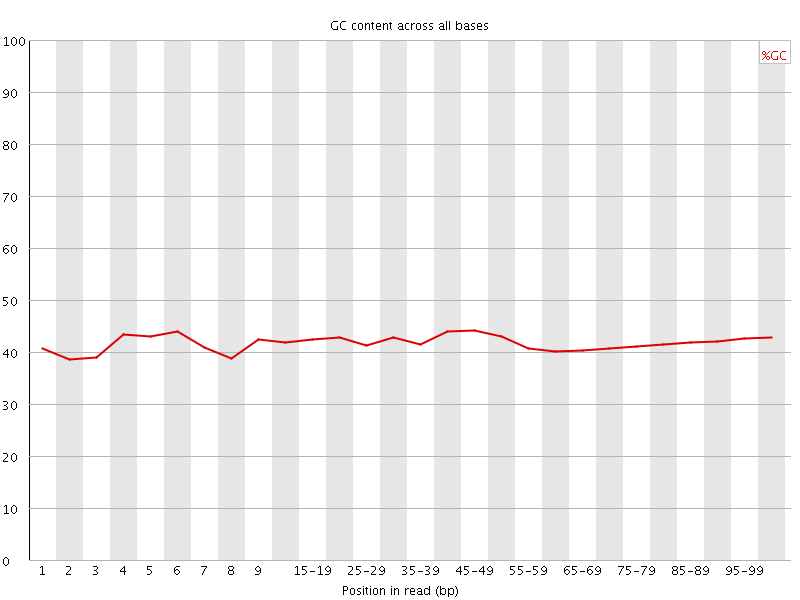
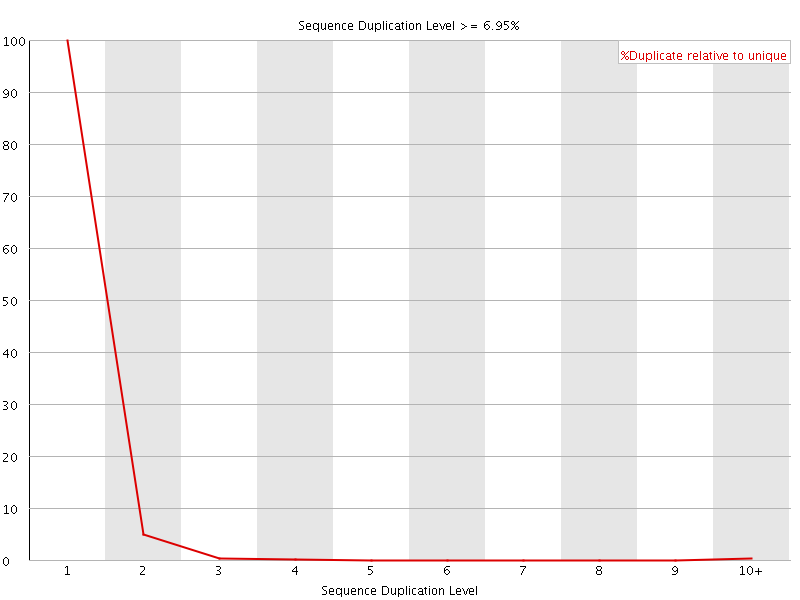
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
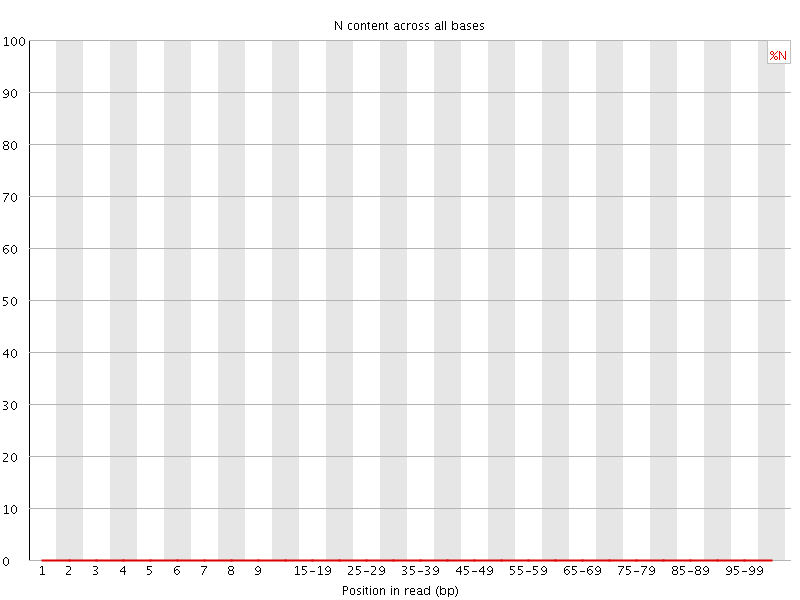
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
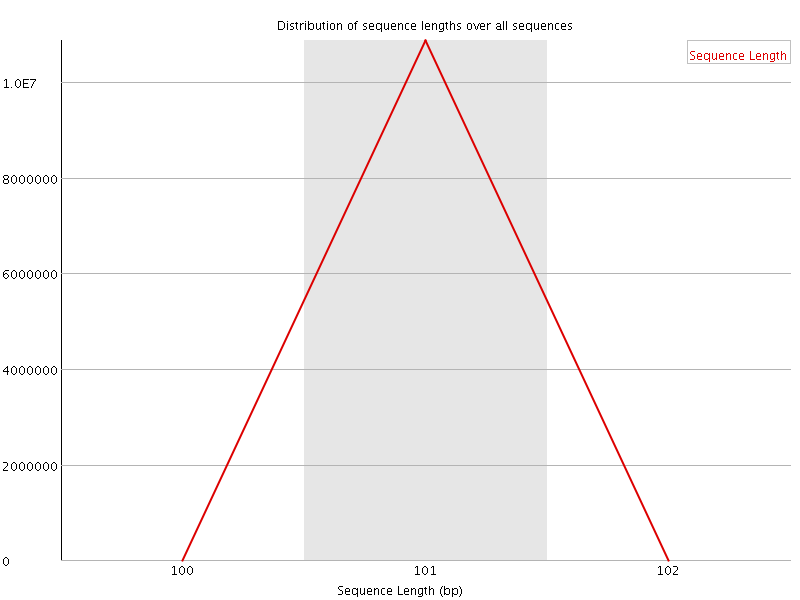
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCTCGGTGGTCGCCG | 14002 | 0.1288250159375185 | Illumina Single End PCR Primer 1 (100% over 50bp) |
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGGGTAGATCTCGGGGGGCGCCG | 11351 | 0.10443456334143496 | Illumina Single End PCR Primer 1 (97% over 41bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
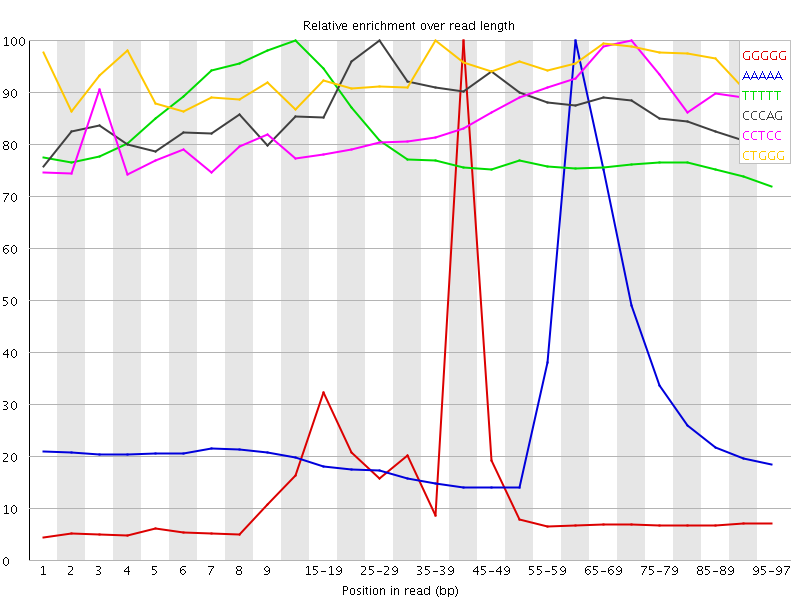
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 3589830 | 6.9643245 | 43.276684 | 40-44 |
| AAAAA | 13267250 | 5.275164 | 18.307089 | 60-64 |
| TTTTT | 6792130 | 3.7009552 | 4.6348476 | 10-14 |
| CCCAG | 2039250 | 3.5696552 | 4.0629 | 25-29 |
| CCTCC | 1763165 | 3.5304186 | 4.0999827 | 70-74 |
| CTGGG | 1991055 | 3.2180722 | 3.4176378 | 35-39 |
| CTCCC | 1574460 | 3.152571 | 3.644278 | 70-74 |
| CCTGG | 1742425 | 3.0246391 | 3.2613466 | 65-69 |
| GGGAG | 2123605 | 3.0006053 | 21.486511 | 5 |
| GGAGG | 2096685 | 2.9625678 | 5.048752 | 6 |
| GGAAG | 2588885 | 2.6642678 | 27.08777 | 5 |
| GGGGC | 1209900 | 2.520934 | 10.058053 | 40-44 |
| GGGGA | 1771045 | 2.502446 | 9.9415045 | 20-24 |
| AGGGG | 1688880 | 2.3863487 | 8.188073 | 25-29 |
| GAGGG | 1613670 | 2.280079 | 12.68889 | 9 |
| AAGAG | 2831320 | 2.1221888 | 19.49365 | 7 |
| GGGAA | 2040430 | 2.099843 | 10.7274 | 20-24 |
| GAAGA | 2672740 | 2.003327 | 19.849913 | 6 |
| AGAGC | 1794765 | 1.9837165 | 33.51906 | 8 |
| GGAGA | 1916430 | 1.9722323 | 16.080143 | 6 |
| AGAGG | 1833325 | 1.8867074 | 9.33456 | 8 |
| GGGCC | 828310 | 1.853582 | 5.9925528 | 45-49 |
| AGGGA | 1785810 | 1.837809 | 5.115785 | 20-24 |
| GGAAA | 2377770 | 1.7822348 | 5.748259 | 20-24 |
| GAGAG | 1686105 | 1.7352008 | 15.383193 | 7 |
| AGATC | 1946445 | 1.6688399 | 6.6574316 | 95-97 |
| GATCG | 1351175 | 1.5905747 | 40.18491 | 1 |
| CGGGG | 739000 | 1.5397722 | 8.898511 | 35-39 |
| ATCGG | 1258840 | 1.4818801 | 40.11911 | 2 |
| GAGCG | 896985 | 1.3612163 | 44.65569 | 9 |
| GGGCG | 635325 | 1.3237561 | 5.9343925 | 40-44 |
| GGCGG | 622445 | 1.2969195 | 5.9892163 | 10-14 |
| TCGGA | 1079570 | 1.2708473 | 28.204916 | 3 |
| CGGAA | 1090815 | 1.2056552 | 27.578213 | 4 |
| TAGGG | 1063155 | 1.1652852 | 6.4895205 | 15-19 |
| GCGTC | 660920 | 1.1472772 | 7.4979033 | 10-14 |
| GTAGG | 1001910 | 1.0981567 | 6.473681 | 15-19 |
| GCGGG | 498545 | 1.0387628 | 5.6826215 | 10-14 |
| CGTCG | 544905 | 0.9458892 | 7.362695 | 10-14 |
| GCCGG | 407155 | 0.91112643 | 6.7664227 | 45-49 |
| TCGGG | 562145 | 0.90857524 | 18.015707 | 3 |
| CTCGG | 510425 | 0.8860361 | 6.2609386 | 35-39 |
| AGCGT | 734075 | 0.86413777 | 5.1053395 | 10-14 |
| GGCGC | 379100 | 0.8483454 | 5.247972 | 40-44 |
| GCGCC | 348255 | 0.8369956 | 5.5540757 | 45-49 |
| CGTGT | 665355 | 0.834193 | 5.7104206 | 15-19 |
| CGGGA | 445590 | 0.67620355 | 19.480185 | 4 |
| CGCCG | 236460 | 0.56830764 | 6.938456 | 45-49 |