![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10869007 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
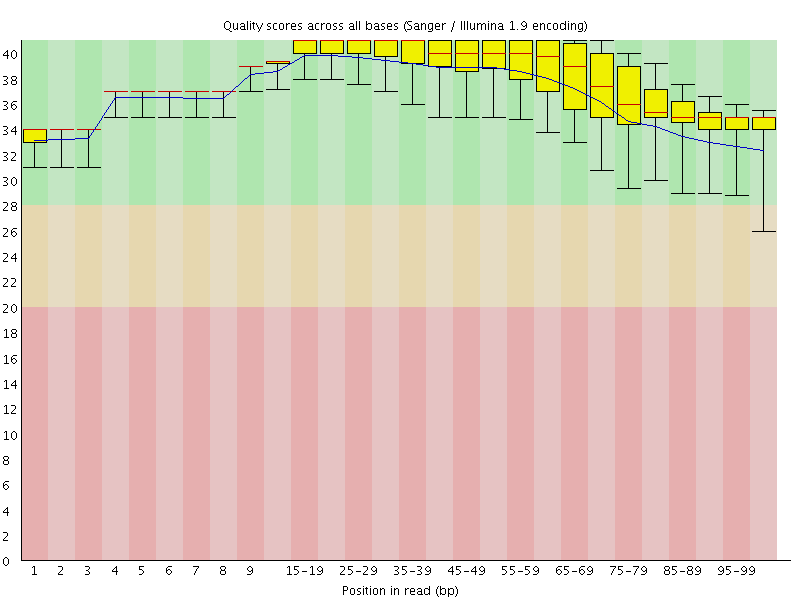
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
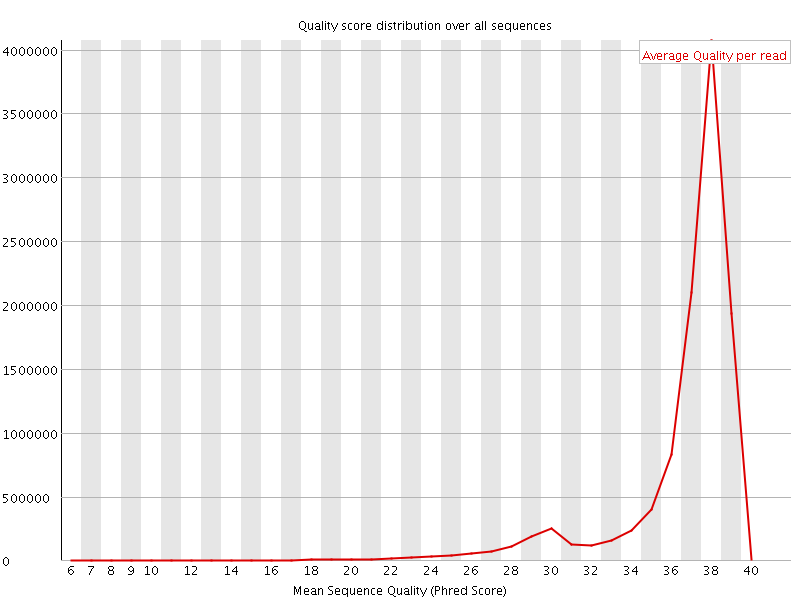
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
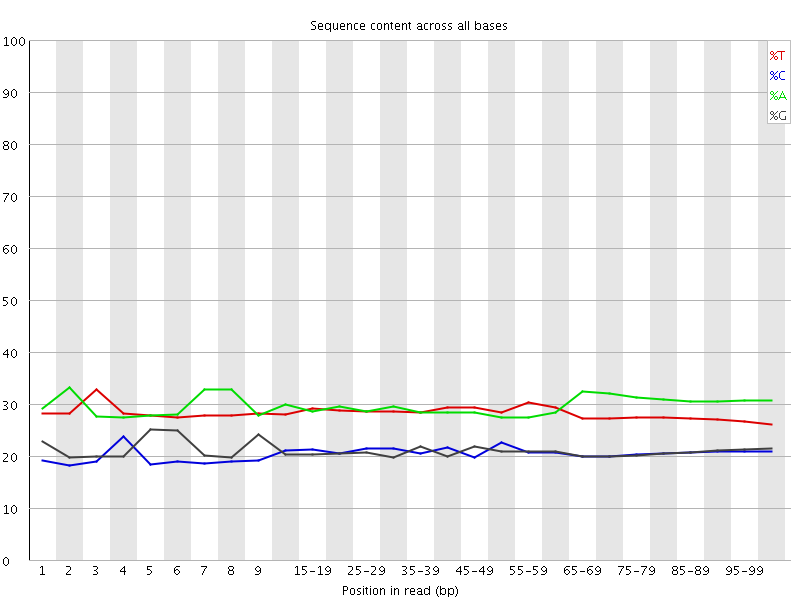
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
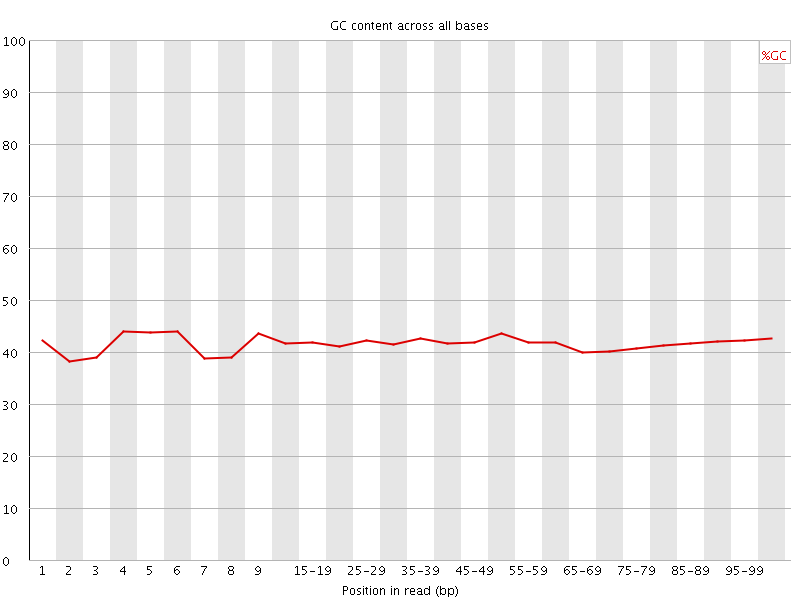
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
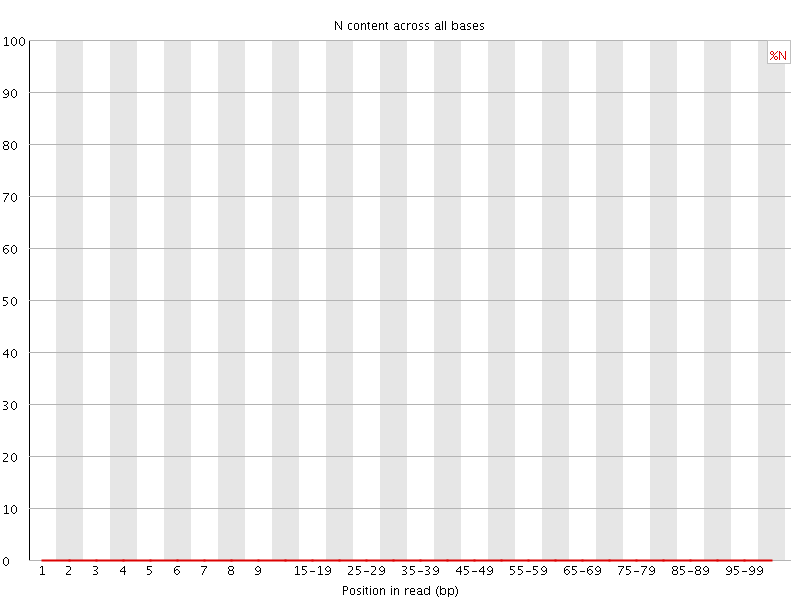
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
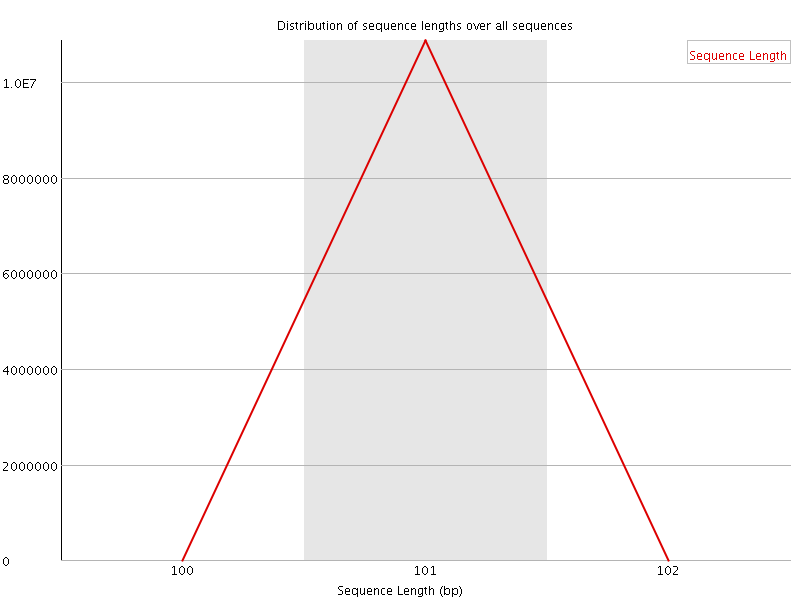
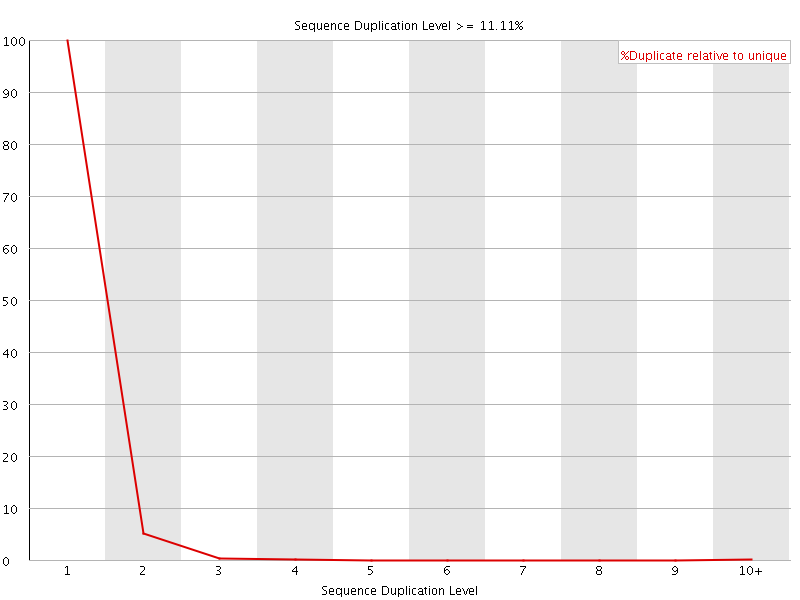
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGC | 477524 | 4.39344642983485 | TruSeq Adapter, Index 5 (100% over 50bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
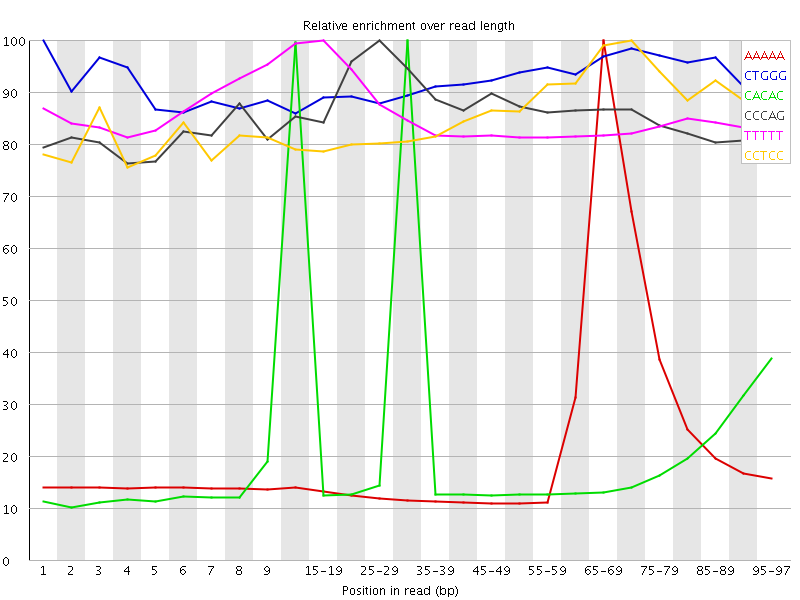
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 12626380 | 5.1180778 | 21.879183 | 65-69 |
| CTGGG | 1984235 | 3.4585724 | 3.7417414 | 1 |
| CACAC | 2909830 | 3.427267 | 13.795973 | 30-34 |
| CCCAG | 2034170 | 3.4071524 | 3.934768 | 25-29 |
| TTTTT | 6561250 | 3.3592737 | 3.9205124 | 15-19 |
| CCTCC | 1774460 | 3.1250007 | 3.6066837 | 70-74 |
| CCTGG | 1753530 | 3.0669758 | 3.296629 | 70-74 |
| GGAAG | 2626890 | 3.06226 | 61.69405 | 5 |
| GGAGG | 1840440 | 3.0510259 | 3.4609761 | 4 |
| CCAGG | 1812565 | 3.0255516 | 3.4297068 | 25-29 |
| CTCCA | 2099565 | 2.5911694 | 14.507771 | 20-24 |
| CAGTG | 2095515 | 2.5684462 | 14.379246 | 35-39 |
| TCCAG | 2050035 | 2.521357 | 14.192118 | 25-29 |
| CACAG | 2083020 | 2.4450085 | 14.001747 | 30-34 |
| CTGCT | 1890370 | 2.4361615 | 15.018838 | 55-59 |
| AGAGC | 2010390 | 2.3516562 | 60.94268 | 8 |
| TCTGC | 1810795 | 2.3336115 | 14.915581 | 55-59 |
| TTCTG | 2389130 | 2.2608292 | 11.289177 | 55-59 |
| GAGCA | 1929030 | 2.2564852 | 60.821323 | 9 |
| GCACA | 1903240 | 2.2339861 | 13.244396 | 10-14 |
| GAAGA | 2720210 | 2.2298646 | 43.38034 | 6 |
| AGCAC | 1889875 | 2.2182987 | 13.119656 | 10-14 |
| AAGAG | 2661285 | 2.1815617 | 43.28973 | 7 |
| CTTCT | 2250530 | 2.1370082 | 11.471727 | 50-54 |
| ACACA | 2578005 | 2.1278777 | 10.0407 | 30-34 |
| ACTCC | 1677505 | 2.0702858 | 14.102466 | 20-24 |
| GTCTG | 1599780 | 2.0545945 | 14.330257 | 15-19 |
| TCTTC | 2067135 | 1.962864 | 11.353002 | 50-54 |
| CGTCT | 1502890 | 1.9368075 | 13.488655 | 15-19 |
| CCAGT | 1565775 | 1.9257611 | 13.774099 | 25-29 |
| TGCTT | 1993170 | 1.886133 | 11.301661 | 55-59 |
| GAAAA | 3246090 | 1.8711672 | 7.450244 | 60-64 |
| GATCG | 1525985 | 1.8703805 | 63.31105 | 1 |
| TCTGA | 2057800 | 1.8584249 | 10.673142 | 15-19 |
| CTGAA | 2139520 | 1.8440472 | 10.176448 | 15-19 |
| ATGCC | 1471920 | 1.810328 | 13.78502 | 45-49 |
| GCTTG | 1397775 | 1.79516 | 14.429228 | 55-59 |
| AGTGA | 2026550 | 1.7406826 | 9.980137 | 35-39 |
| ATCGG | 1403345 | 1.7200623 | 63.256344 | 2 |
| TCACA | 1985615 | 1.7172917 | 9.949882 | 30-34 |
| CAGTC | 1379220 | 1.6963155 | 13.554235 | 25-29 |
| TCGGA | 1356995 | 1.6632516 | 63.20224 | 3 |
| TGAAA | 2725010 | 1.645911 | 7.4650903 | 60-64 |
| ATCTC | 1804695 | 1.6354564 | 10.109447 | 40-44 |
| CTTGA | 1809685 | 1.6343491 | 10.36236 | 60-64 |
| CGGAA | 1350910 | 1.5802288 | 60.331158 | 4 |
| GTCTT | 1661330 | 1.5721134 | 10.804132 | 50-54 |
| GTCAC | 1272085 | 1.5645491 | 13.614001 | 25-29 |
| AGATC | 1797390 | 1.5491658 | 6.5494123 | 95-97 |
| AACTC | 1715200 | 1.4834188 | 9.819832 | 20-24 |
| TGAAC | 1708965 | 1.4729527 | 9.650281 | 20-24 |
| GTGAT | 1632395 | 1.4691755 | 10.094617 | 35-39 |
| TTGAA | 2305090 | 1.4588555 | 7.6264997 | 60-64 |
| ACAGT | 1688205 | 1.4550599 | 9.841981 | 30-34 |
| GAACT | 1645840 | 1.4185452 | 9.686728 | 20-24 |
| AGTCA | 1608795 | 1.3866165 | 9.868408 | 25-29 |
| TGATC | 1455150 | 1.3141639 | 9.952772 | 35-39 |
| CCGTC | 732145 | 1.2849542 | 17.955198 | 50-54 |
| CACGT | 1044625 | 1.284794 | 13.078234 | 10-14 |
| GATCT | 1421365 | 1.2836524 | 9.903889 | 35-39 |
| TGCCG | 707405 | 1.2372723 | 18.592226 | 45-49 |
| ACACG | 1047685 | 1.2297524 | 12.478316 | 10-14 |
| GCCGT | 669885 | 1.1716486 | 18.267666 | 45-49 |
| TATGC | 1224285 | 1.1056669 | 9.865459 | 45-49 |
| GTATG | 1221120 | 1.099023 | 9.71129 | 45-49 |
| ACGTC | 886445 | 1.090247 | 12.69372 | 15-19 |
| TCTCG | 841915 | 1.0849944 | 13.330855 | 40-44 |
| CTCGT | 712900 | 0.91872996 | 13.204095 | 40-44 |
| CGTAT | 688010 | 0.6213504 | 9.456603 | 40-44 |
| TCGTA | 649040 | 0.58615613 | 9.5089035 | 40-44 |