![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L008_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 14086255 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
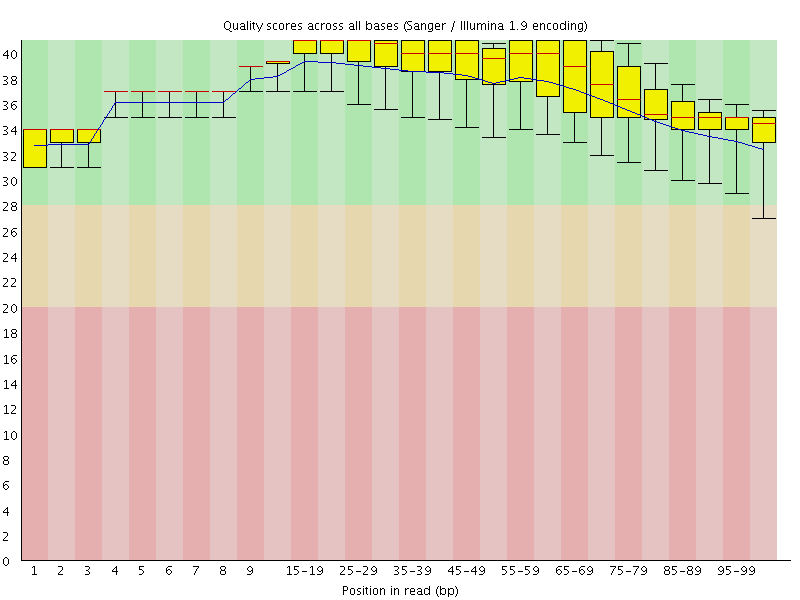
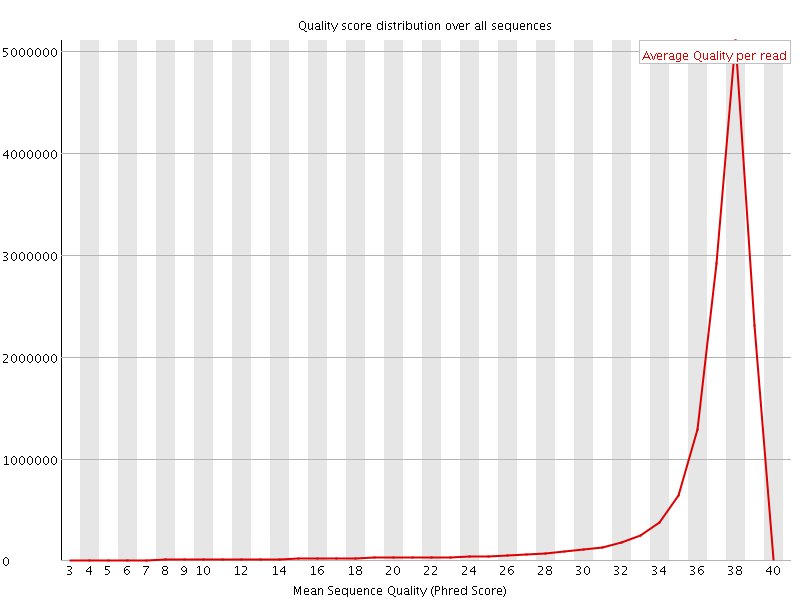
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
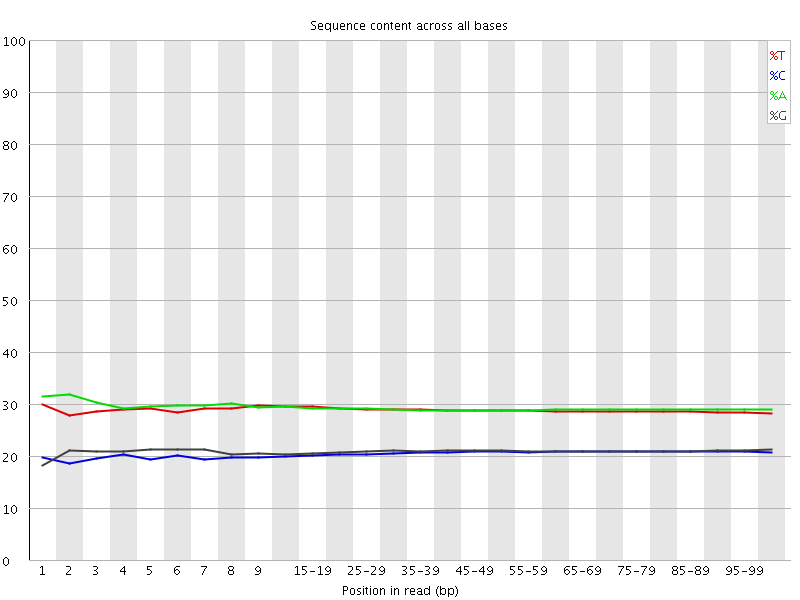
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
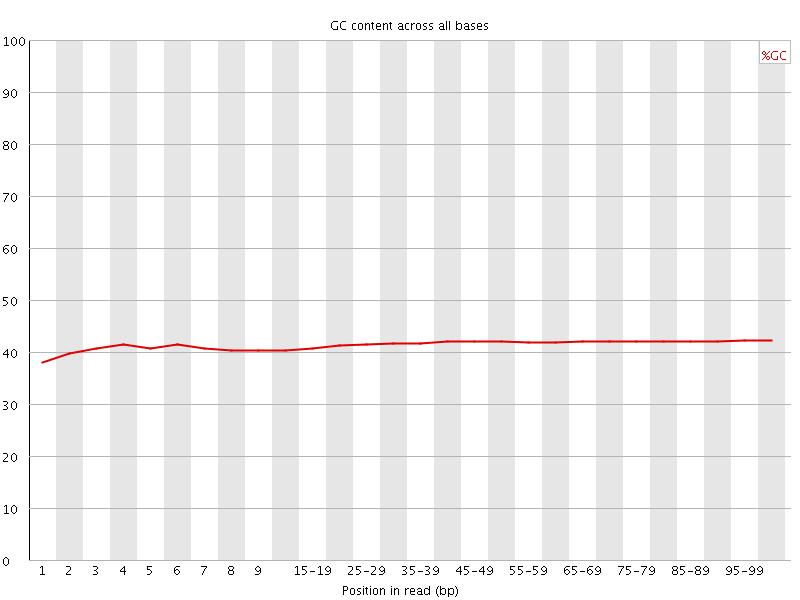
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
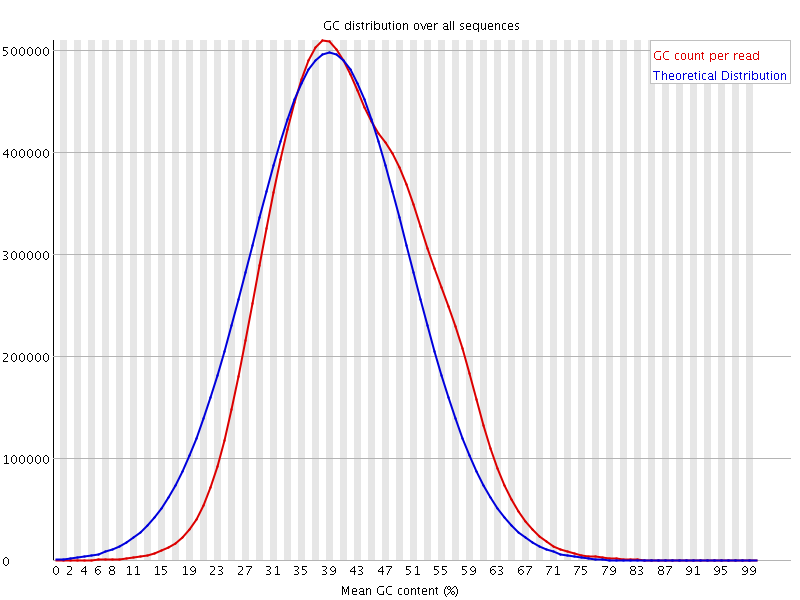
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
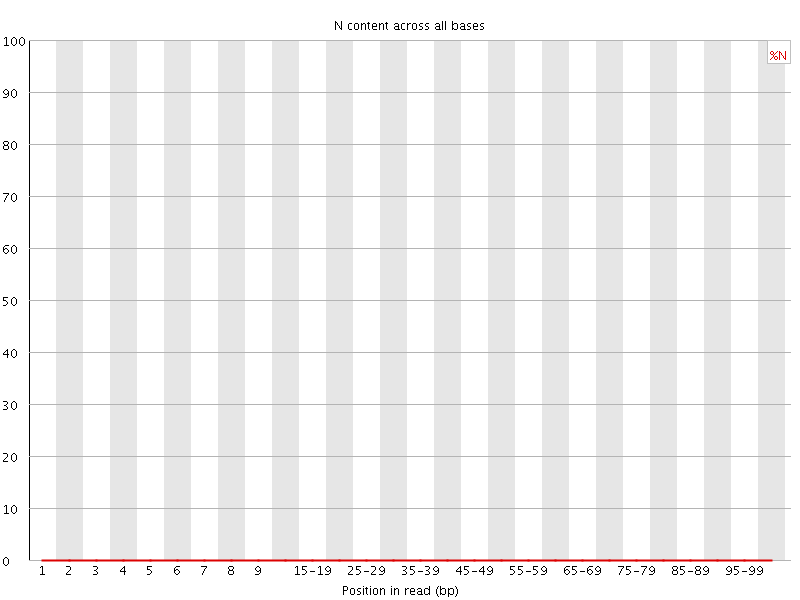
![[OK]](Icons/tick.png) Per base N content
Per base N content
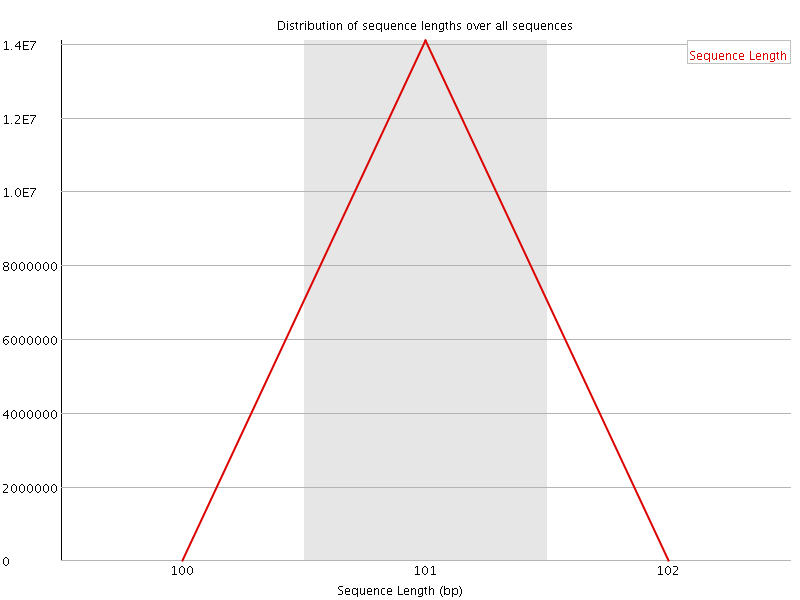
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
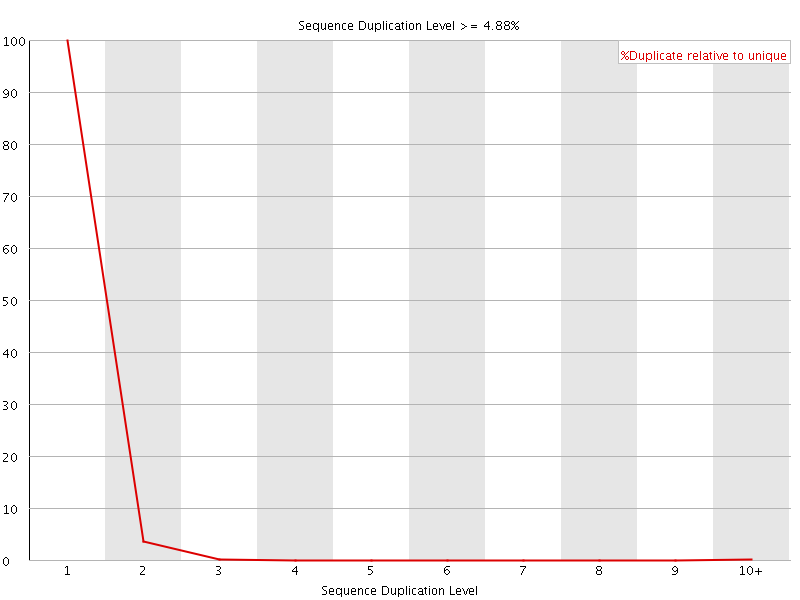
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
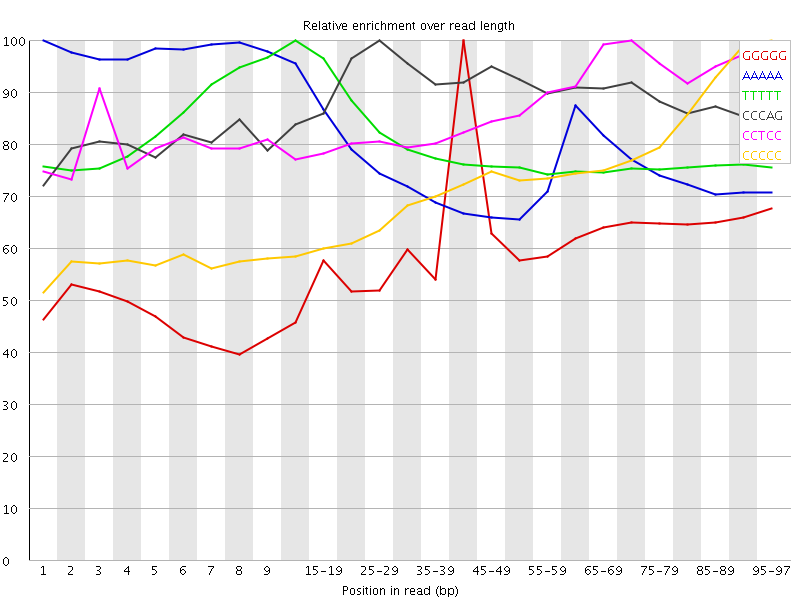
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2323855 | 4.1022177 | 6.765929 | 40-44 |
| AAAAA | 11572645 | 3.9550157 | 5.1125884 | 1 |
| TTTTT | 10411010 | 3.7457268 | 4.684907 | 10-14 |
| CCCAG | 2568260 | 3.428071 | 3.8239522 | 25-29 |
| CCTCC | 2374090 | 3.2542195 | 3.736694 | 70-74 |
| CCCCC | 1698620 | 3.2529135 | 4.438412 | 95-97 |
| GGAGG | 2548750 | 3.2397995 | 3.583336 | 3 |
| CTGGG | 2481130 | 3.2387638 | 3.4404218 | 75-79 |
| CCAGG | 2299320 | 3.0195134 | 3.3655074 | 25-29 |