![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 14086255 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
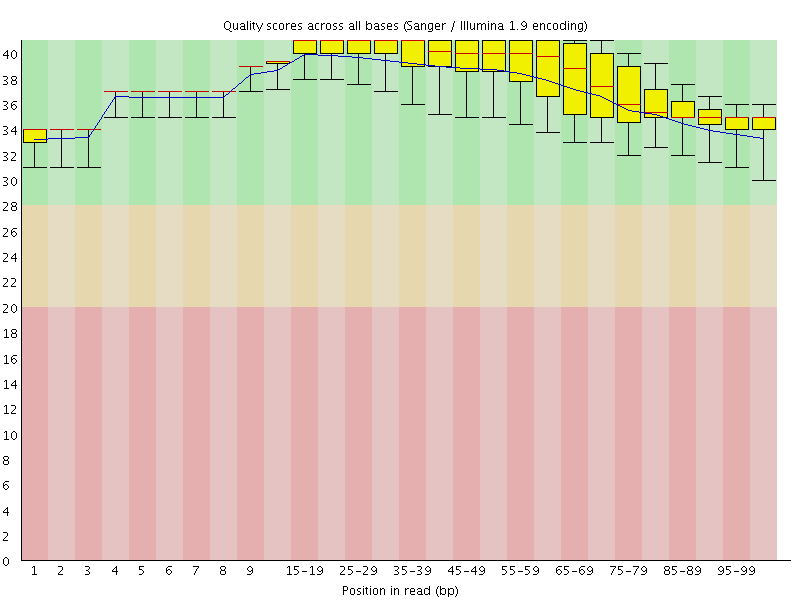
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
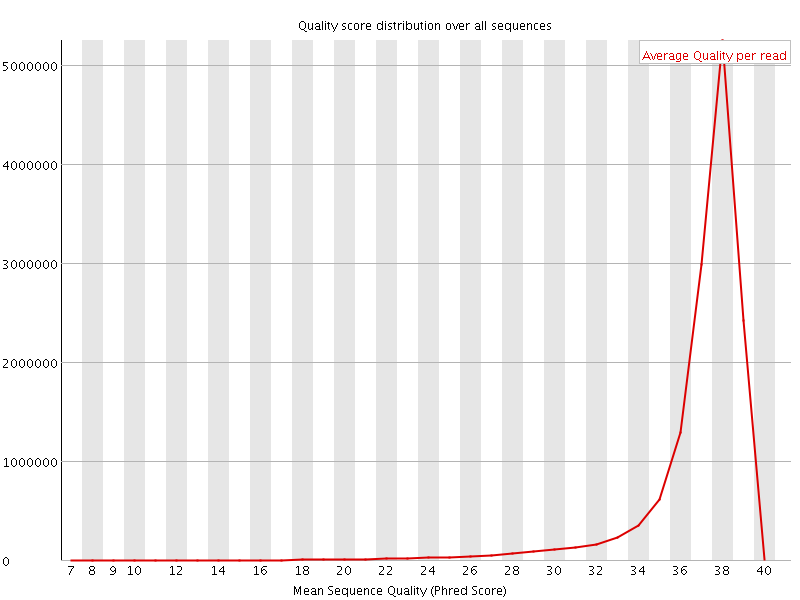
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
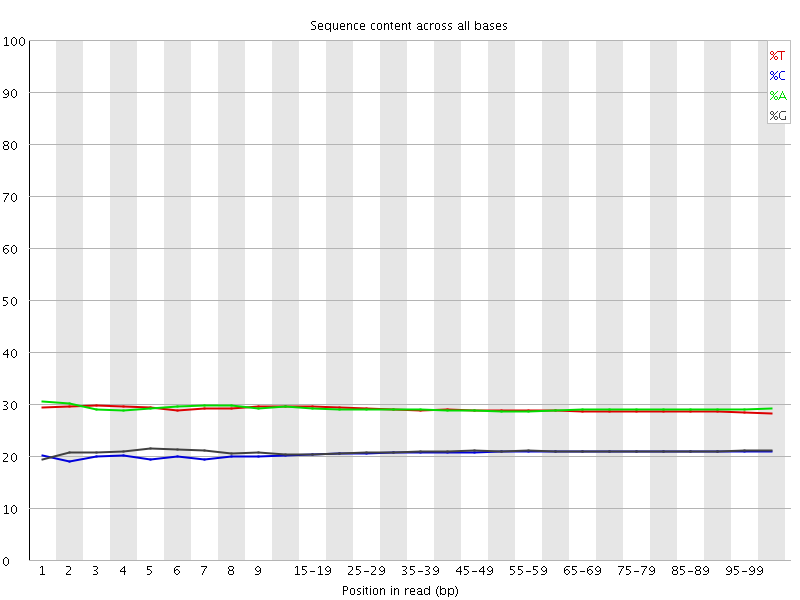
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
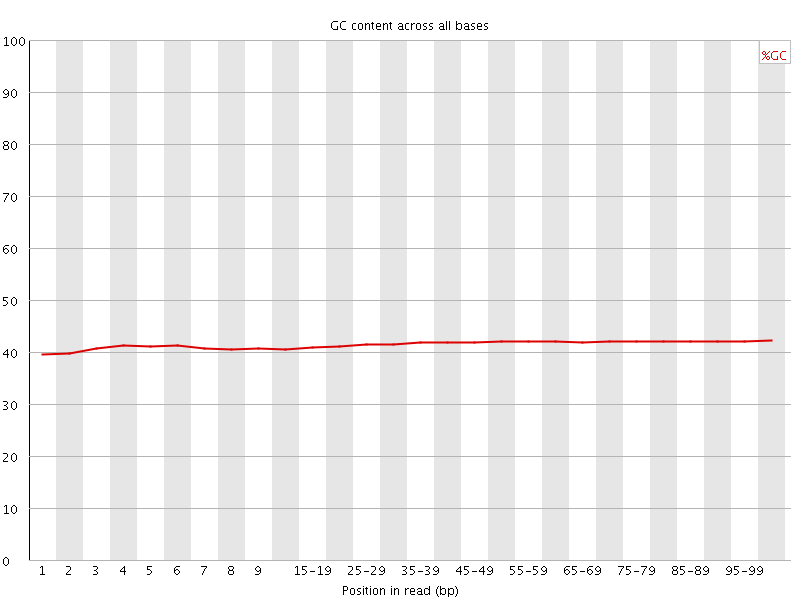
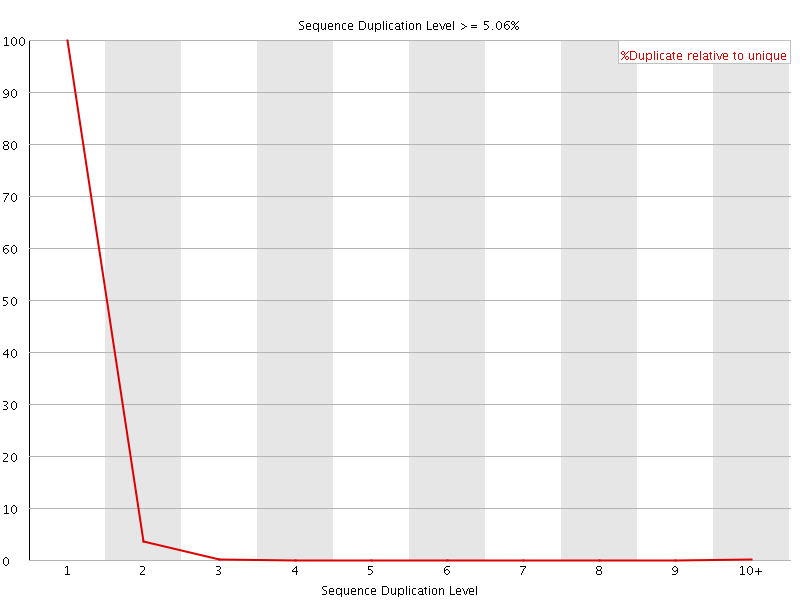
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
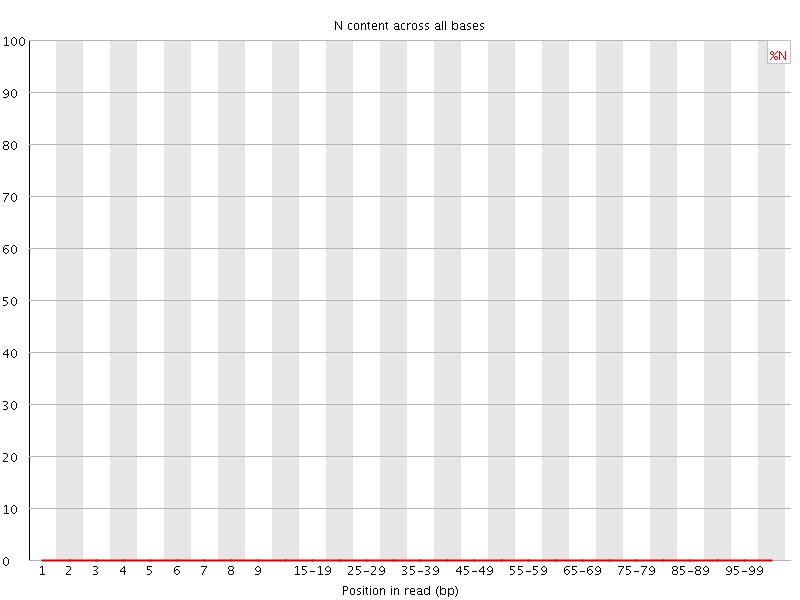
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
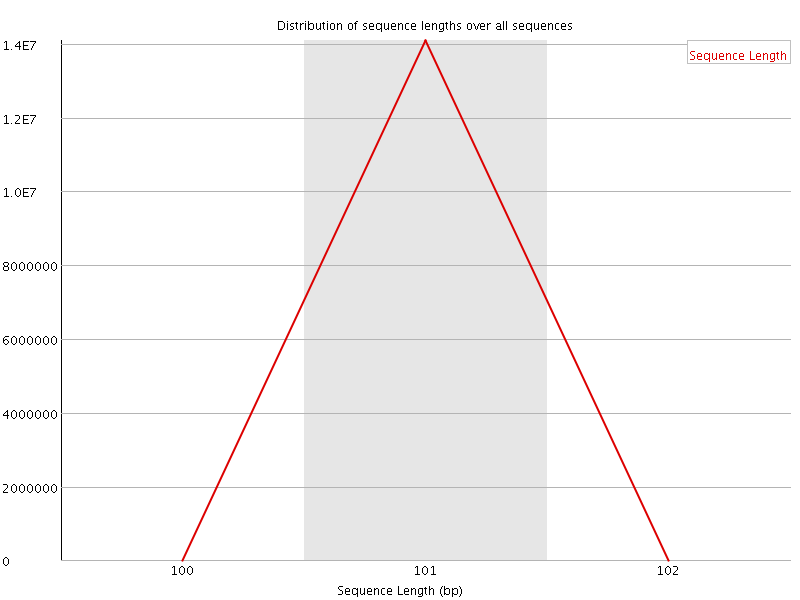
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 38868 | 0.27592855588657167 | TruSeq Adapter, Index 4 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
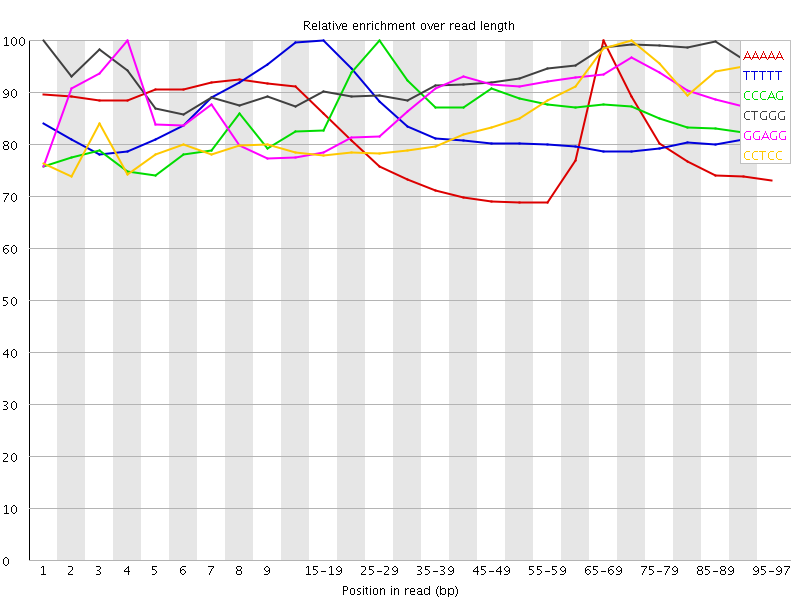
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 10431750 | 3.6380048 | 4.600343 | 65-69 |
| TTTTT | 10239585 | 3.6139174 | 4.3050256 | 15-19 |
| CCCAG | 2570655 | 3.4200063 | 3.9514103 | 25-29 |
| CTGGG | 2491270 | 3.256605 | 3.4744995 | 1 |
| GGAGG | 2510185 | 3.24096 | 3.6662676 | 4 |
| CCTCC | 2391050 | 3.2206838 | 3.7373269 | 70-74 |
| GGGGG | 1746365 | 3.1276517 | 3.816115 | 65-69 |
| CCAGG | 2306820 | 3.038494 | 3.3851418 | 25-29 |
| GGAAG | 2249755 | 2.0940533 | 6.0338893 | 5 |
| GAGCA | 1485975 | 1.3970191 | 5.268714 | 9 |
| AGAGC | 1474665 | 1.3863863 | 5.2170577 | 8 |