![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 14010096 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
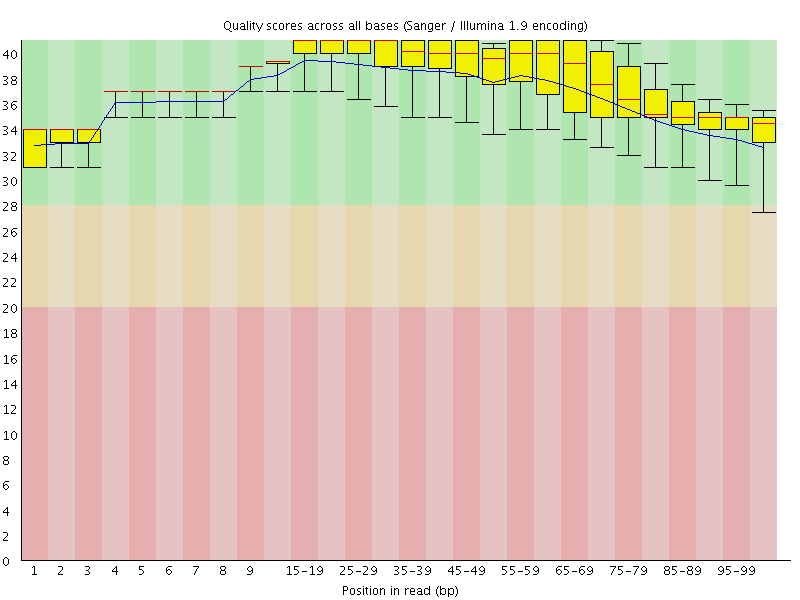
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
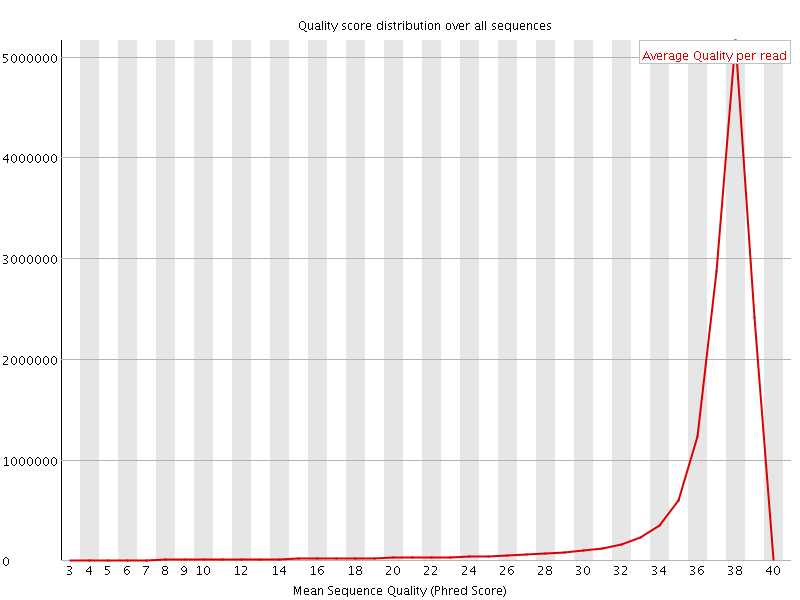
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
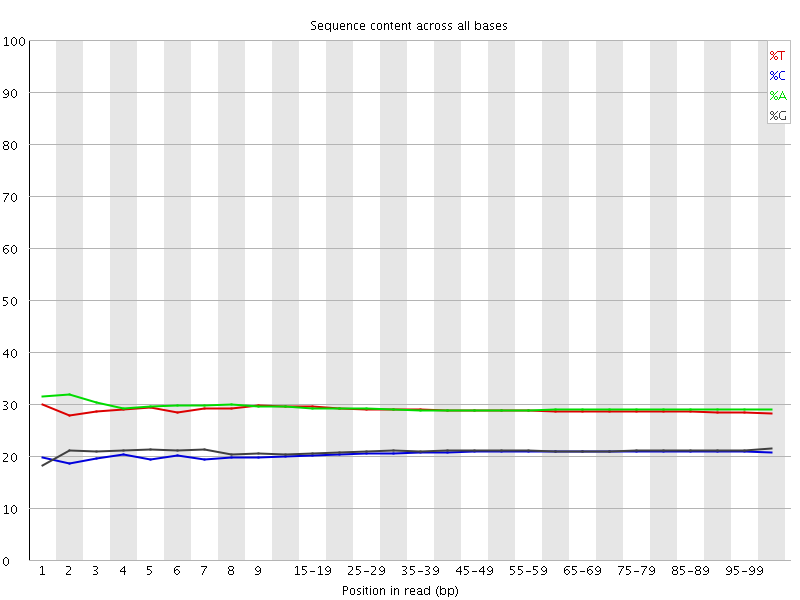
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
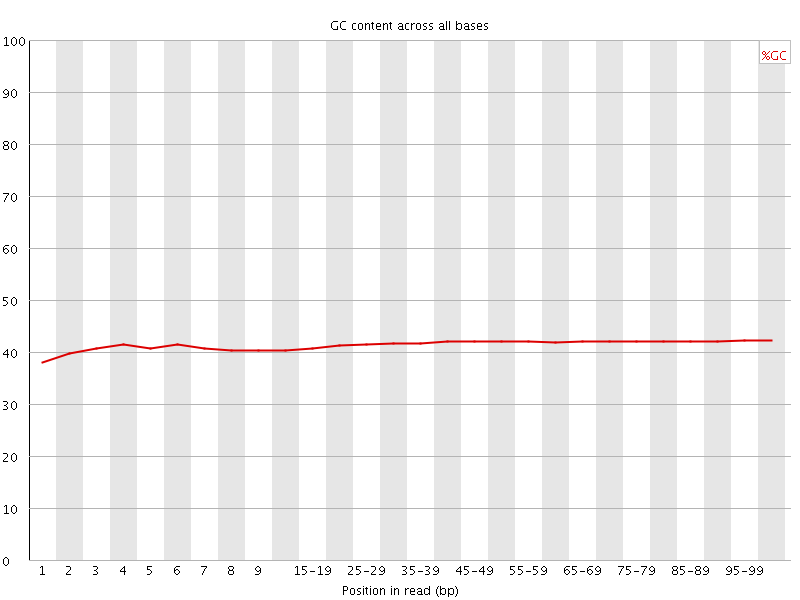
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
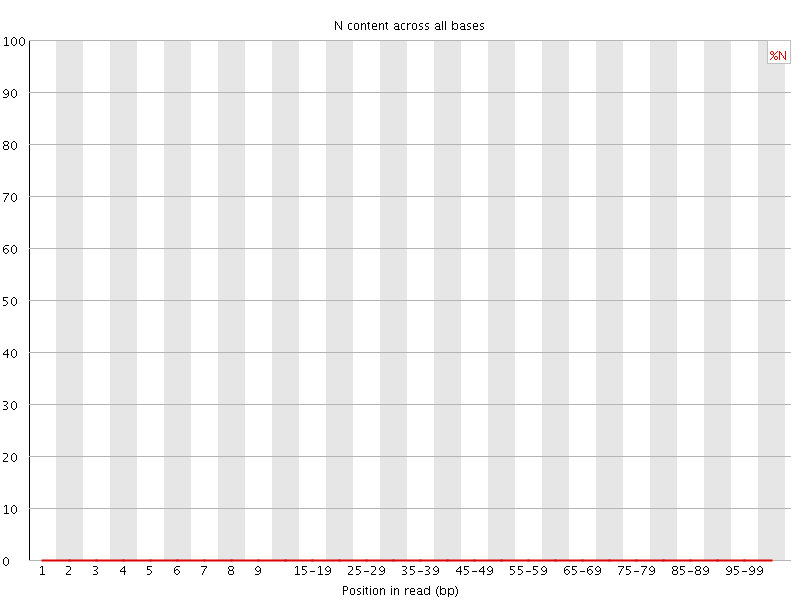
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
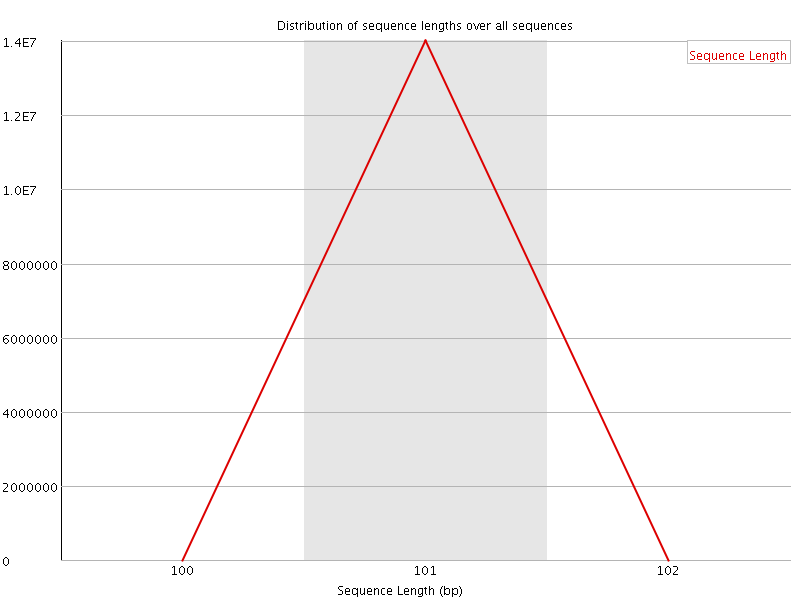
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
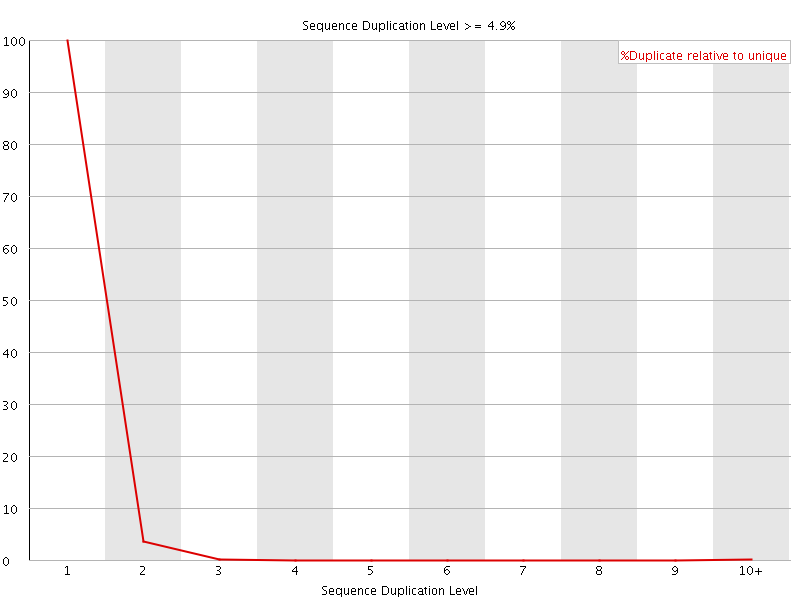
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
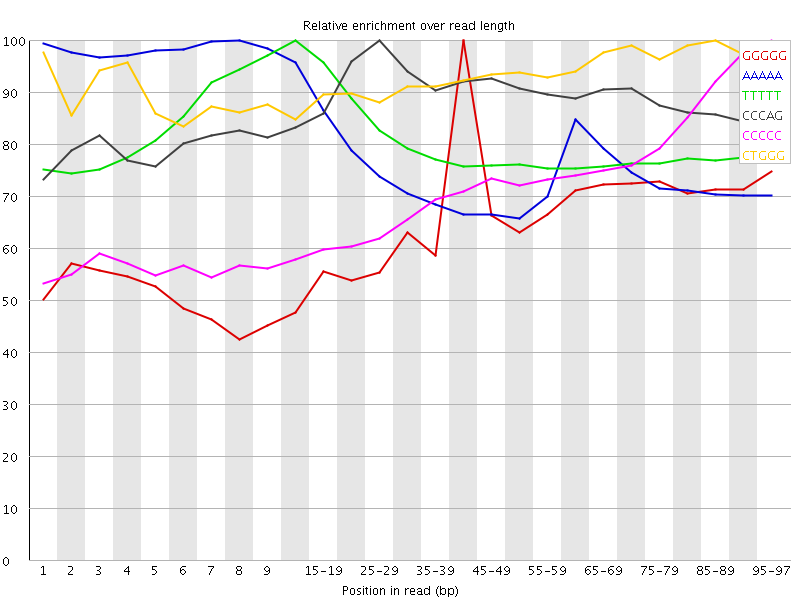
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2277155 | 4.041713 | 6.1775556 | 40-44 |
| AAAAA | 11408380 | 3.9352221 | 5.1377463 | 8 |
| TTTTT | 10435880 | 3.7684116 | 4.684518 | 10-14 |
| CCCAG | 2558330 | 3.428652 | 3.8588276 | 25-29 |
| CCCCC | 1708650 | 3.2781217 | 4.521321 | 95-97 |
| CTGGG | 2471585 | 3.2404072 | 3.4725363 | 85-89 |
| CCTCC | 2357325 | 3.2383437 | 3.7300246 | 70-74 |
| GGAGG | 2528710 | 3.2343454 | 3.6338055 | 3 |
| CCAGG | 2291815 | 3.0240338 | 3.3646812 | 25-29 |