![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 14010096 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
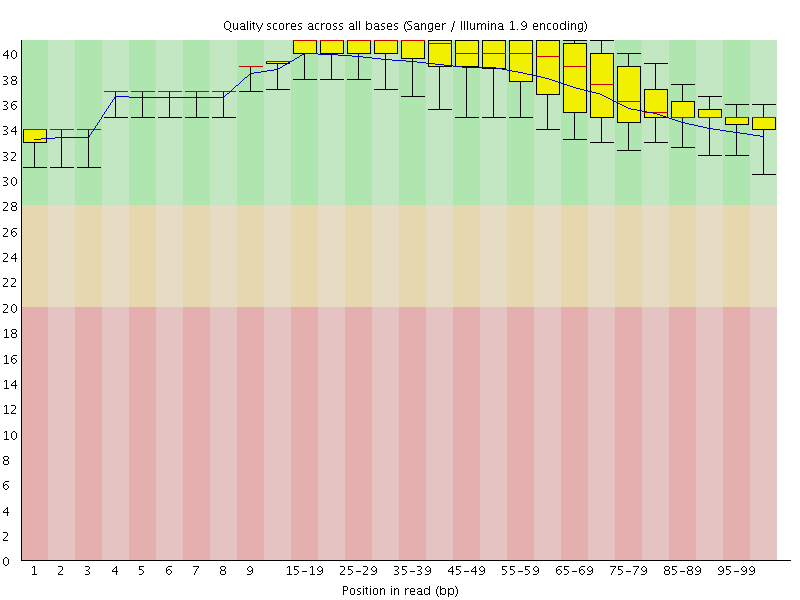
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
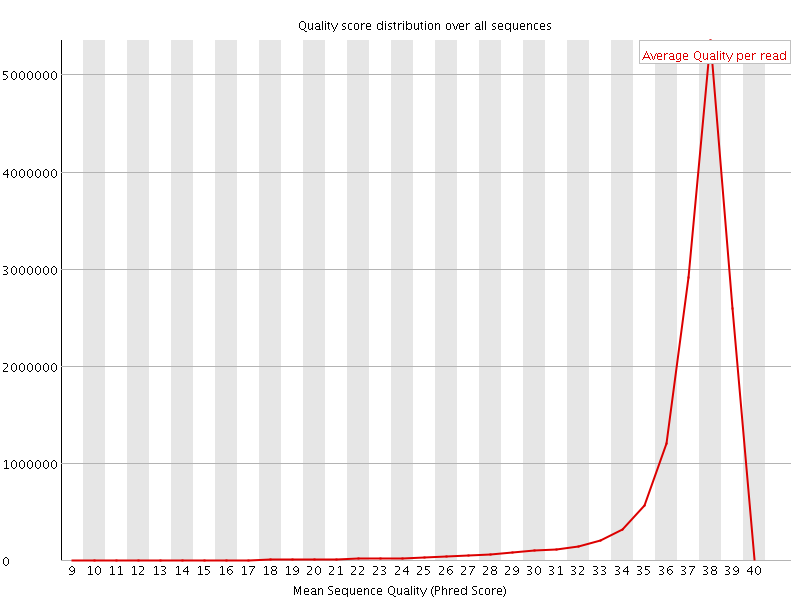
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
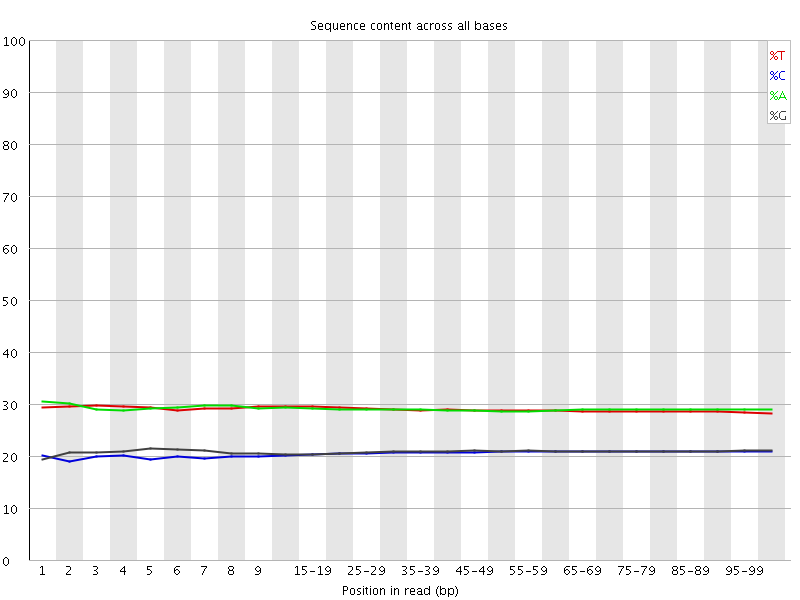
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
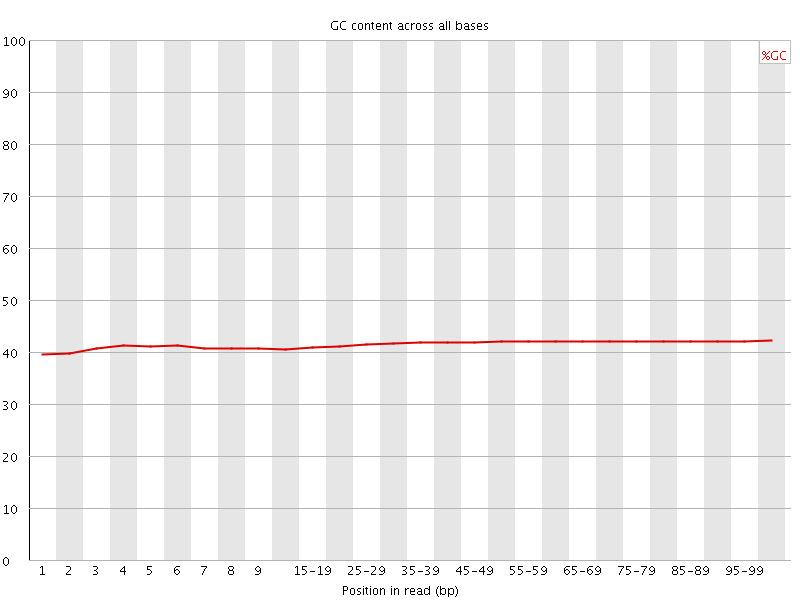
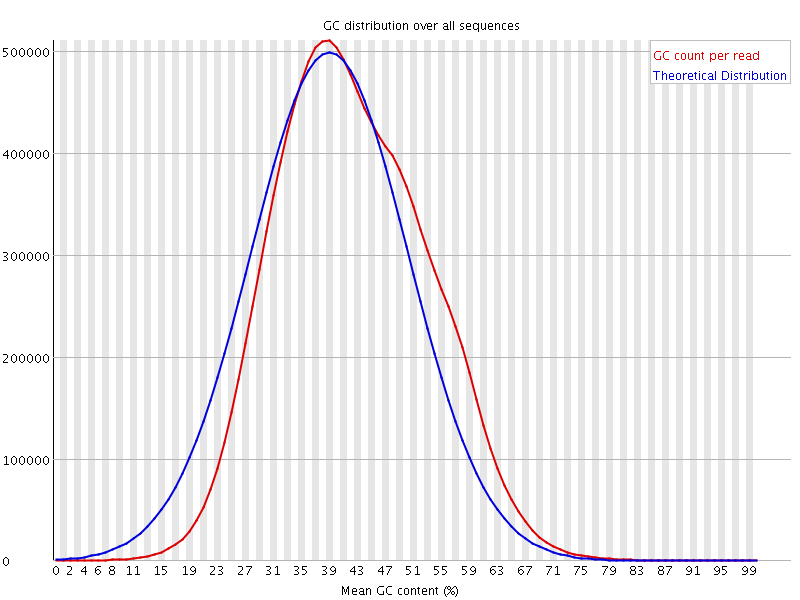
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
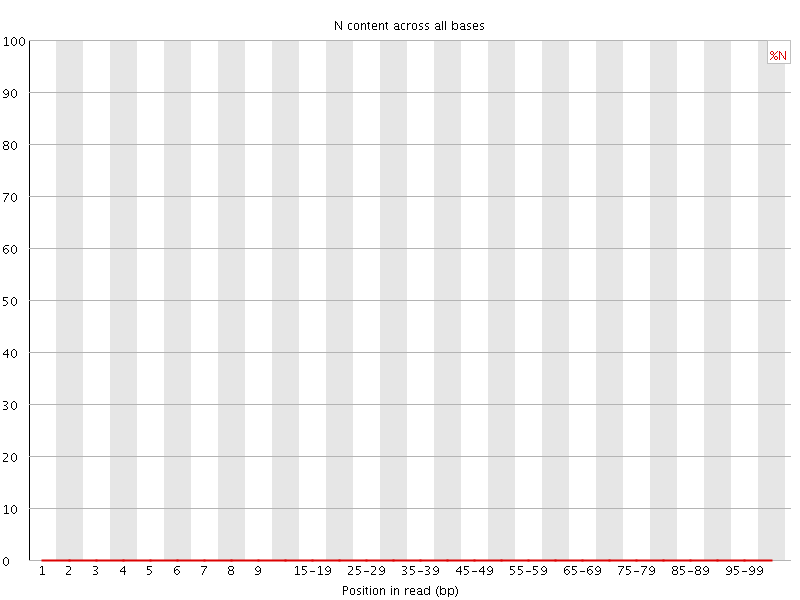
![[OK]](Icons/tick.png) Per base N content
Per base N content
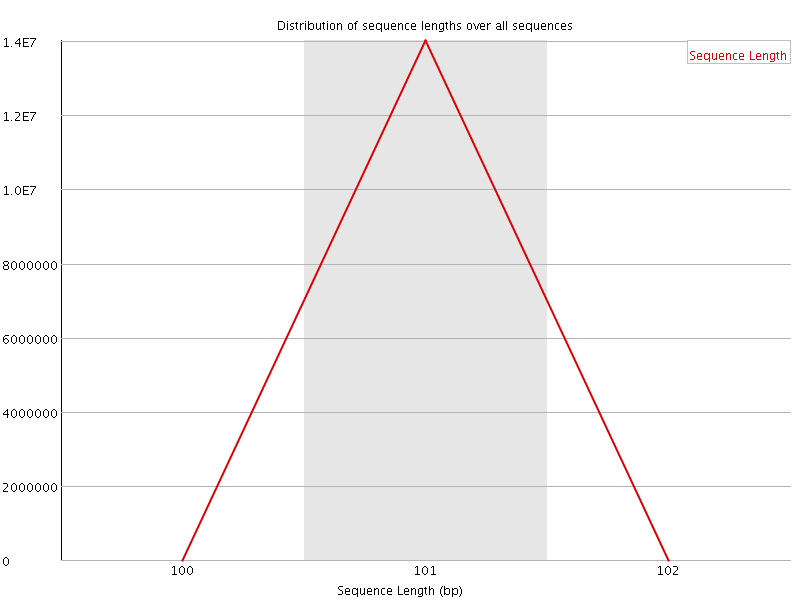
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
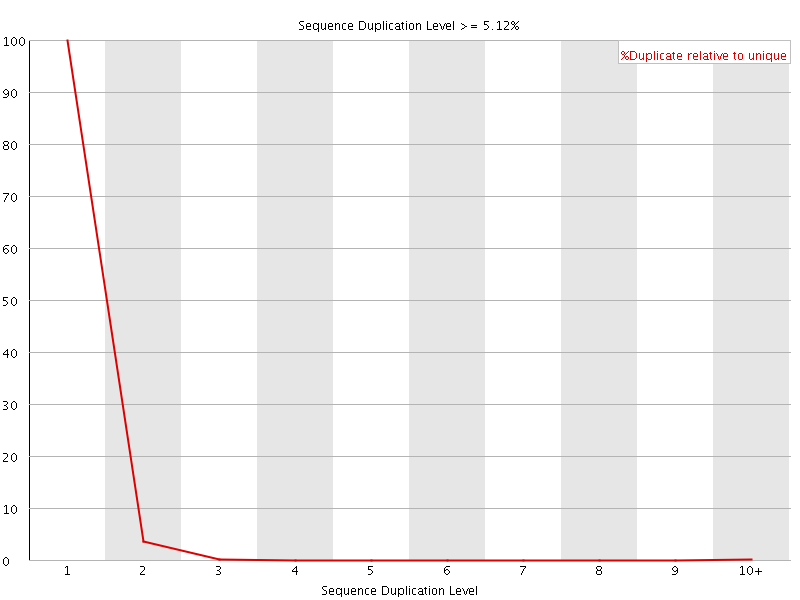
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 34129 | 0.24360289893802298 | TruSeq Adapter, Index 4 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
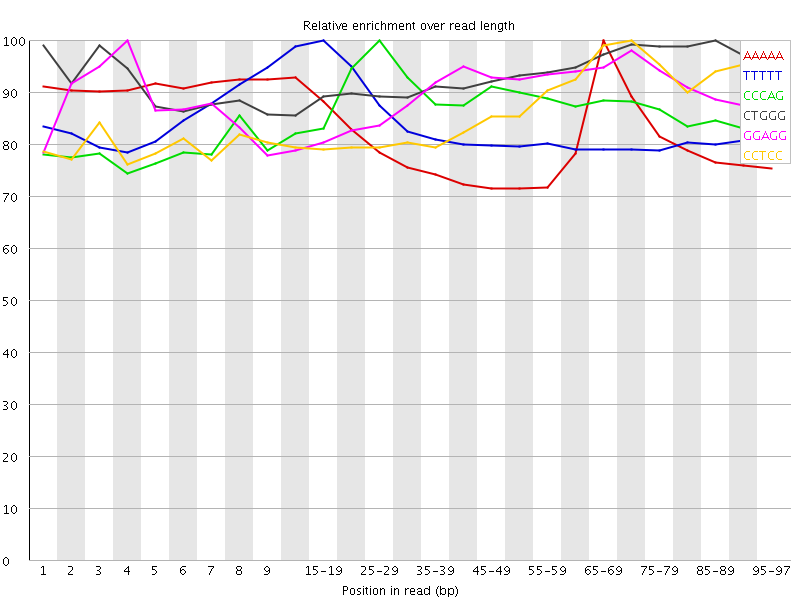
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 10300885 | 3.6209216 | 4.470881 | 65-69 |
| TTTTT | 10090955 | 3.5906093 | 4.2873635 | 15-19 |
| CCCAG | 2562720 | 3.4218805 | 3.9221973 | 25-29 |
| CTGGG | 2481855 | 3.252368 | 3.4797869 | 85-89 |
| GGAGG | 2505740 | 3.2411692 | 3.6161087 | 4 |
| CCTCC | 2371105 | 3.2075396 | 3.6861222 | 70-74 |
| GGGGG | 1743245 | 3.1230536 | 3.8929684 | 65-69 |
| CCAGG | 2301480 | 3.040686 | 3.4259489 | 25-29 |
| GGAAG | 2234295 | 2.0866563 | 5.5533423 | 5 |