![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL1_CGATGT_L008_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 18026677 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
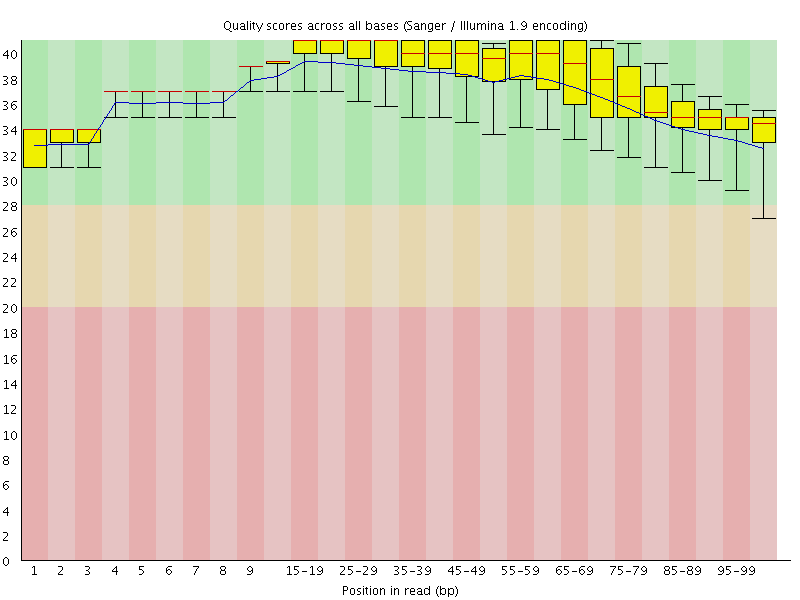
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
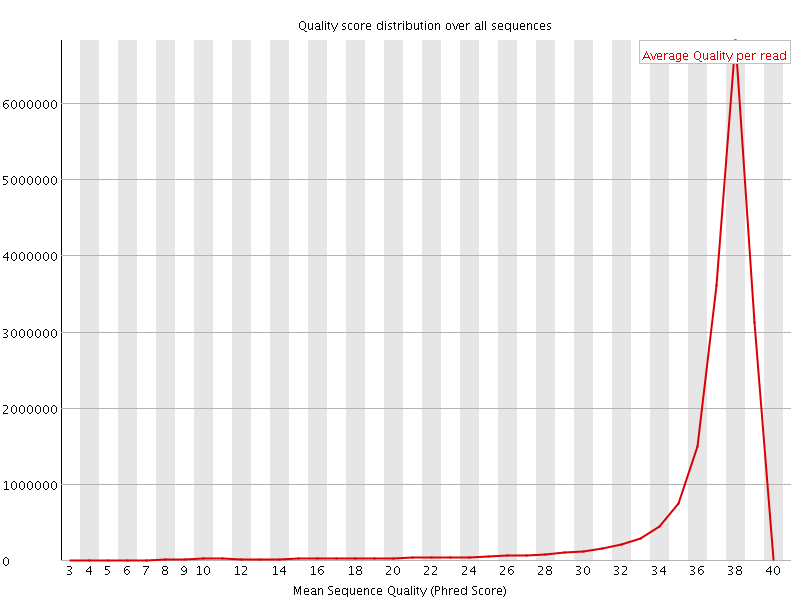
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
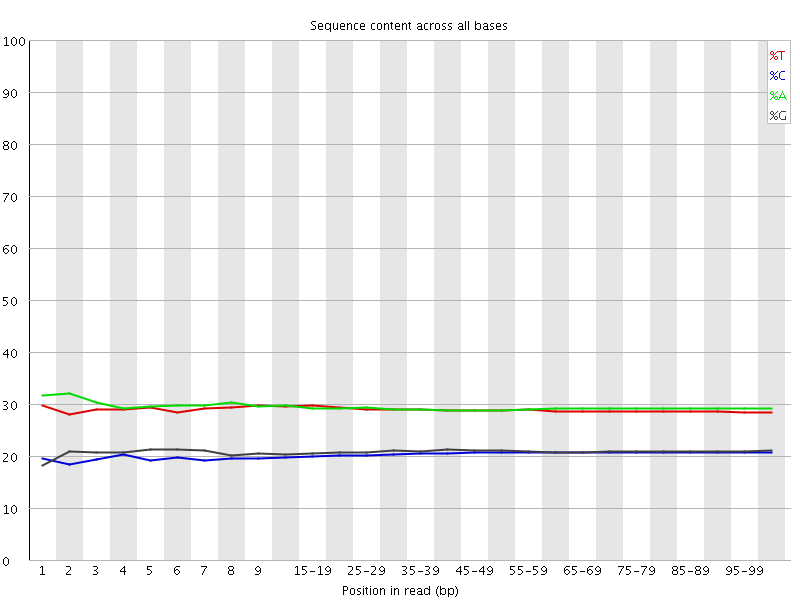
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
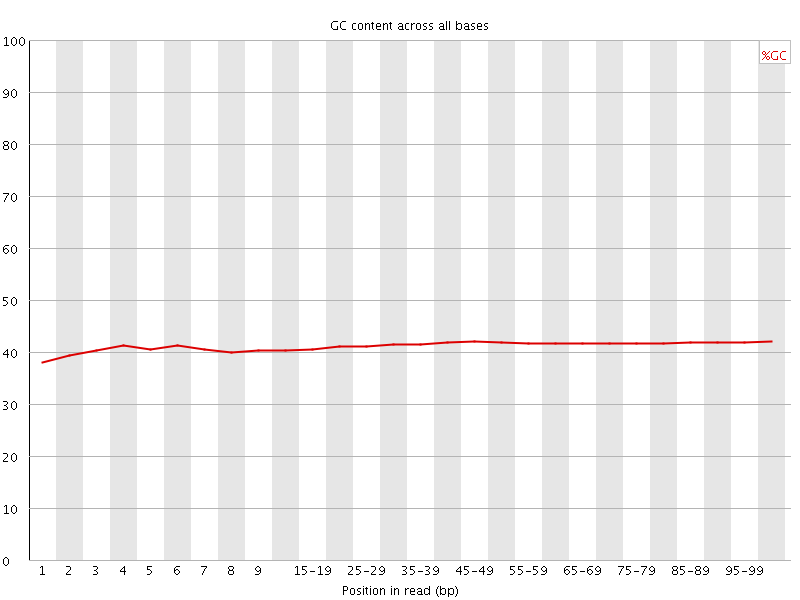
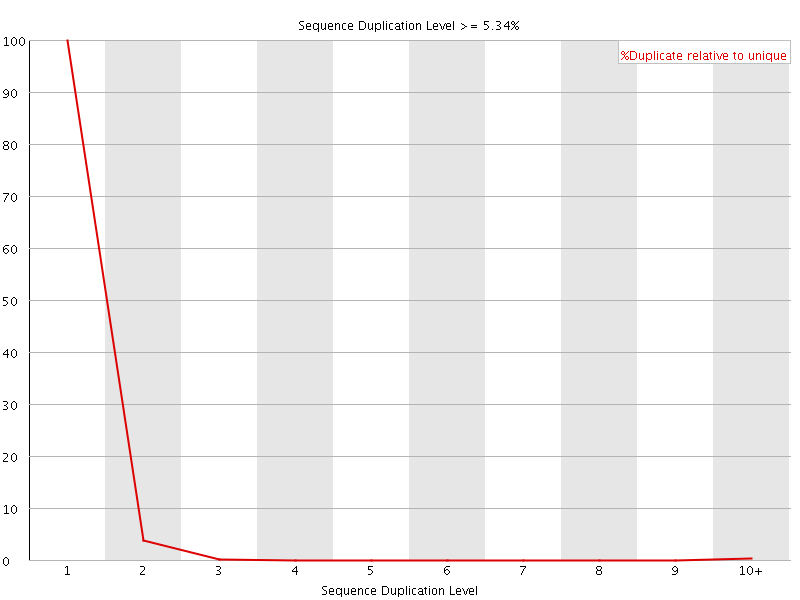
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
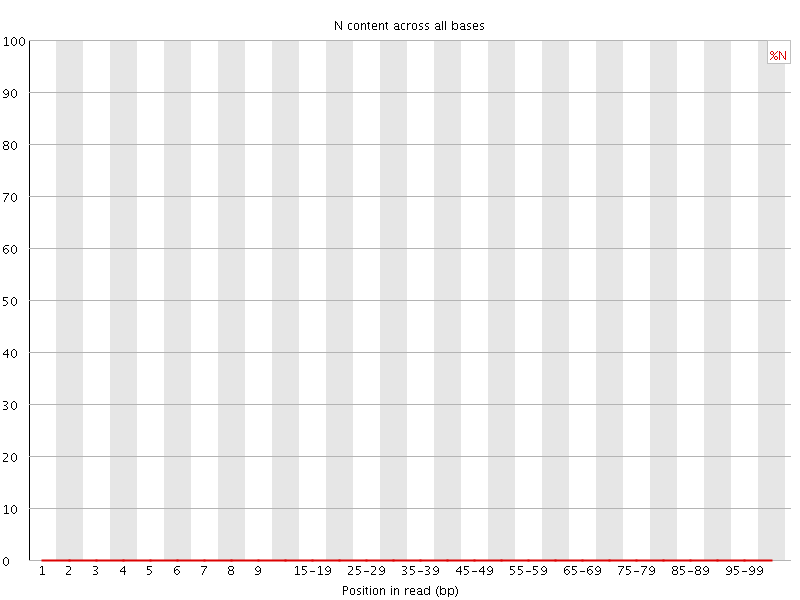
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
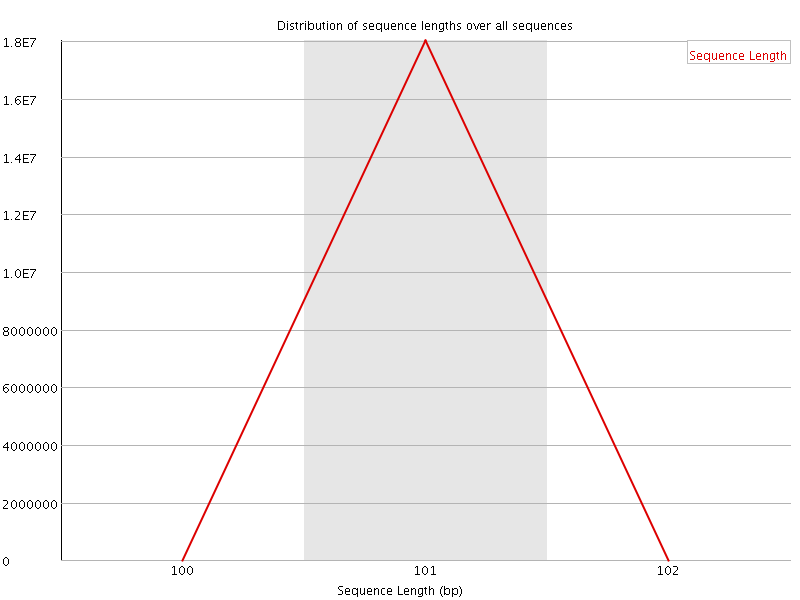
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
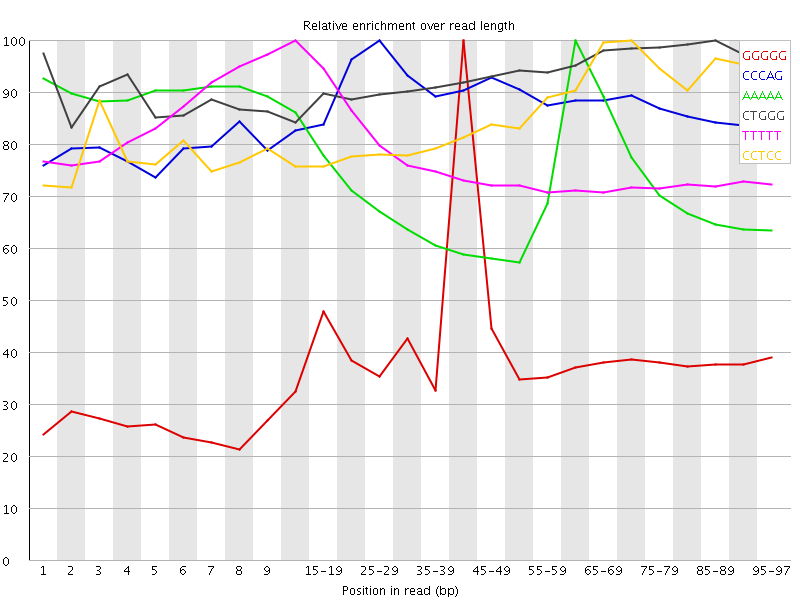
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2817785 | 3.957 | 9.846357 | 40-44 |
| CCCAG | 3489080 | 3.715444 | 4.2251544 | 25-29 |
| AAAAA | 13943890 | 3.6519933 | 5.047635 | 60-64 |
| CTGGG | 3364720 | 3.480863 | 3.7246063 | 85-89 |
| TTTTT | 12103690 | 3.3414516 | 4.3199835 | 10-14 |
| CCTCC | 3007280 | 3.3007753 | 3.8409488 | 70-74 |
| GGAGG | 3279710 | 3.291783 | 3.5688734 | 70-74 |
| CCAGG | 3129445 | 3.2673757 | 3.6640444 | 25-29 |
| CCTGG | 2953390 | 3.1162107 | 3.3669608 | 70-74 |
| GGGAG | 2939280 | 2.9501 | 5.888921 | 5 |
| GGAAG | 2828505 | 2.0290384 | 5.3342776 | 5 |
| AGAGC | 1901335 | 1.391105 | 5.4224644 | 8 |
| GAGCG | 451430 | 0.4621194 | 6.1657395 | 9 |
| GATCG | 540860 | 0.3999083 | 5.131256 | 1 |
| ATCGG | 442865 | 0.32745147 | 5.1110907 | 2 |