![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL1_CGATGT_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 18026677 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
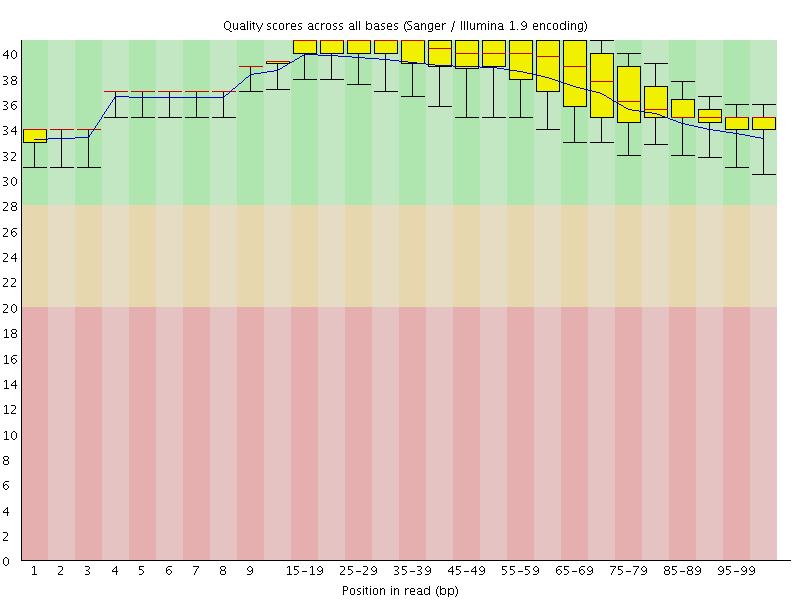
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
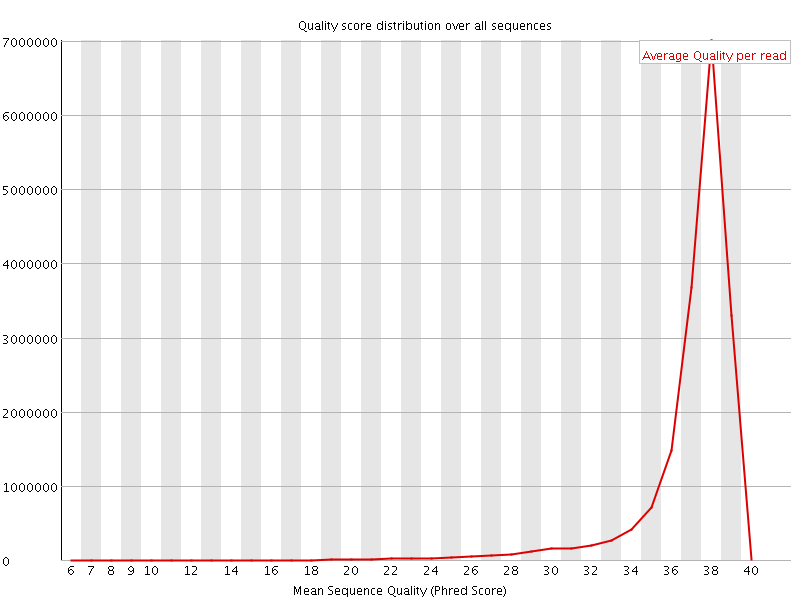
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
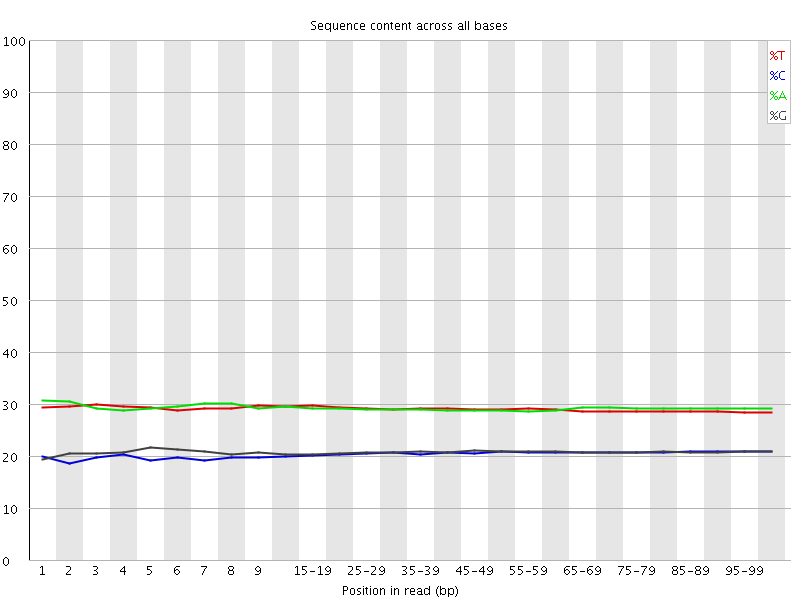
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
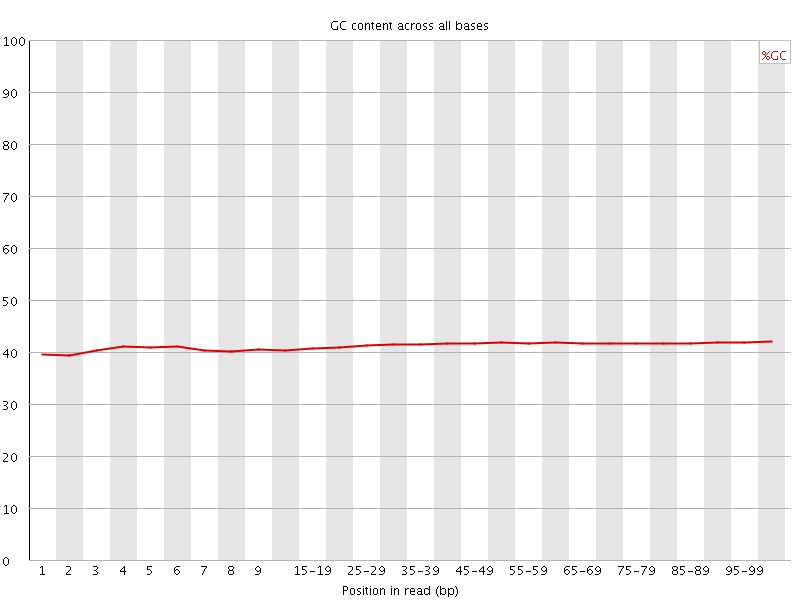
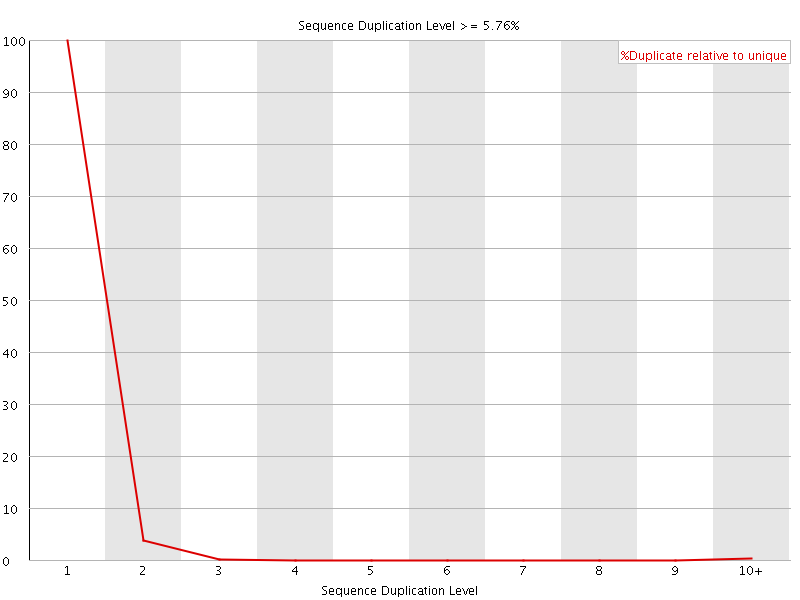
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
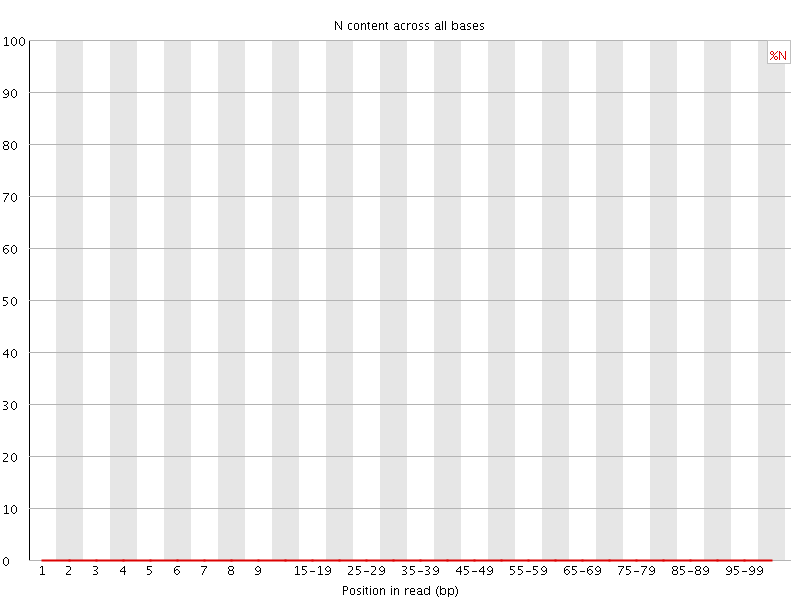
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
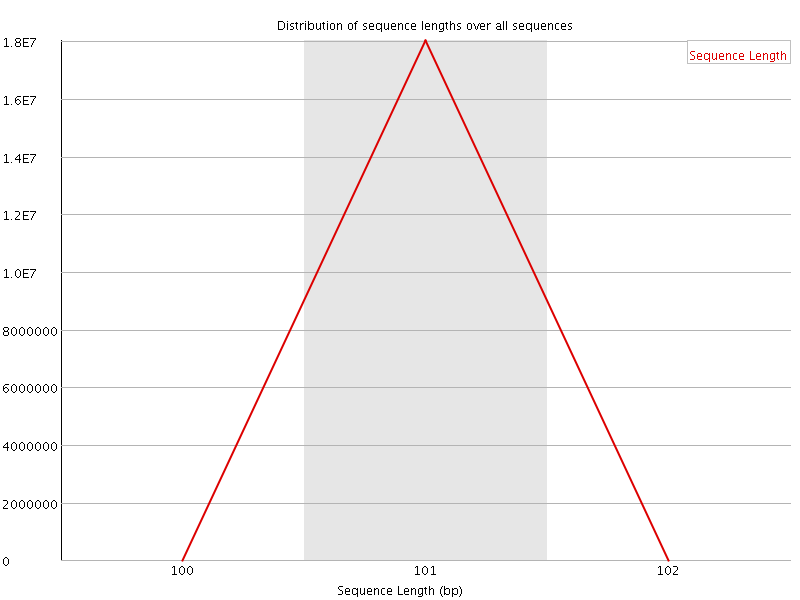
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGC | 105894 | 0.5874293969986815 | TruSeq Adapter, Index 2 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
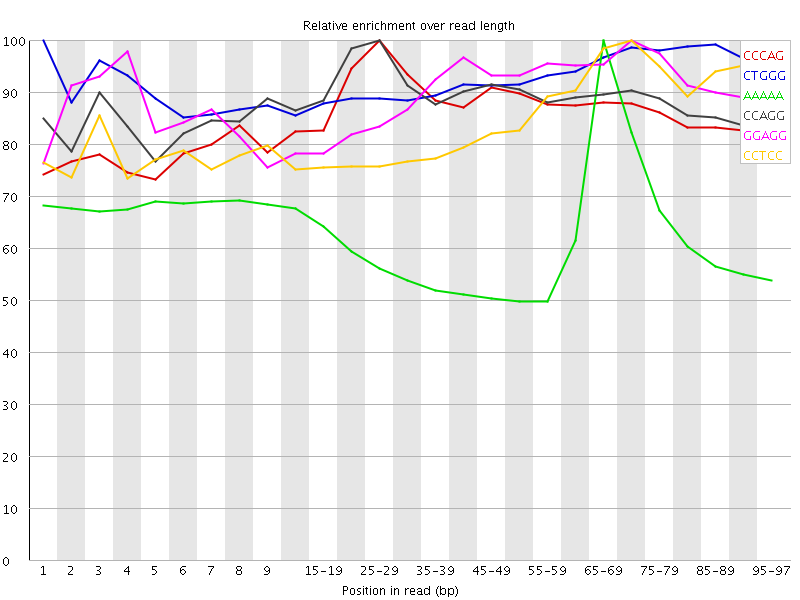
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCCAG | 3506665 | 3.7074013 | 4.2646933 | 25-29 |
| CTGGG | 3378370 | 3.5237617 | 3.7974083 | 1 |
| AAAAA | 12912885 | 3.4282014 | 5.5691395 | 65-69 |
| CCAGG | 3144860 | 3.2948084 | 3.7009623 | 25-29 |
| GGAGG | 3193990 | 3.2860138 | 3.647401 | 70-74 |
| CCTCC | 3023410 | 3.2406776 | 3.8093834 | 70-74 |
| TTTTT | 11757620 | 3.1948285 | 3.8546913 | 10-14 |
| CCTGG | 2963690 | 3.119454 | 3.4029055 | 70-74 |
| GGAAG | 2823665 | 2.0705028 | 10.163666 | 5 |
| GAAGA | 2985910 | 1.5605087 | 7.3205047 | 6 |
| AAGAG | 2973295 | 1.5539156 | 7.2477856 | 7 |
| AGAGC | 1964265 | 1.4534806 | 9.472243 | 8 |
| GAGCA | 1897235 | 1.403881 | 9.438411 | 9 |
| GATCG | 585635 | 0.43536466 | 8.510691 | 1 |
| CGGAA | 554650 | 0.41041967 | 8.5004015 | 4 |
| TCGGA | 492190 | 0.3658971 | 8.500688 | 3 |
| ATCGG | 476955 | 0.35457128 | 8.47807 | 2 |