![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL1_CGATGT_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 17934766 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
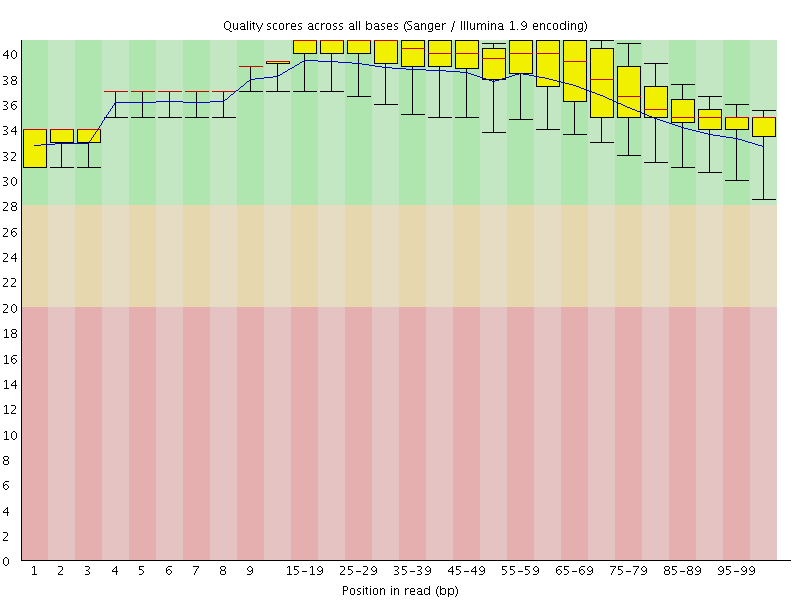
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
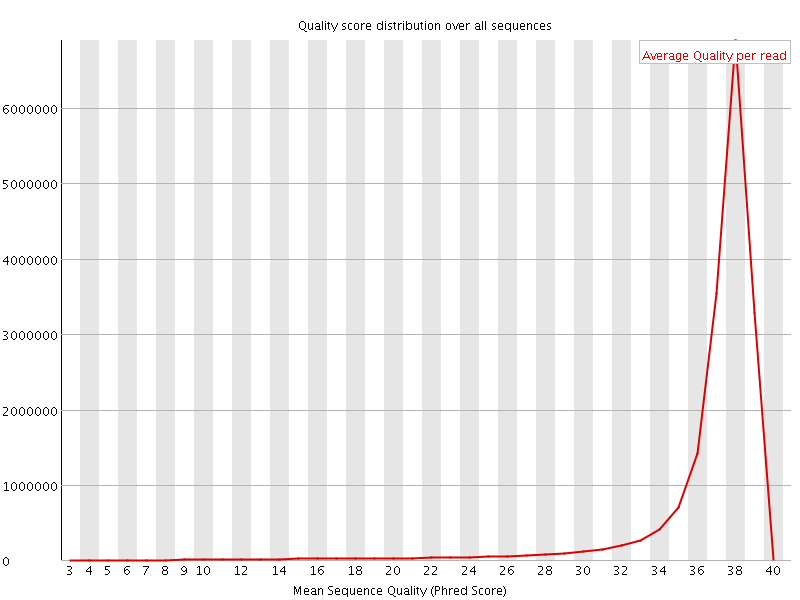
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
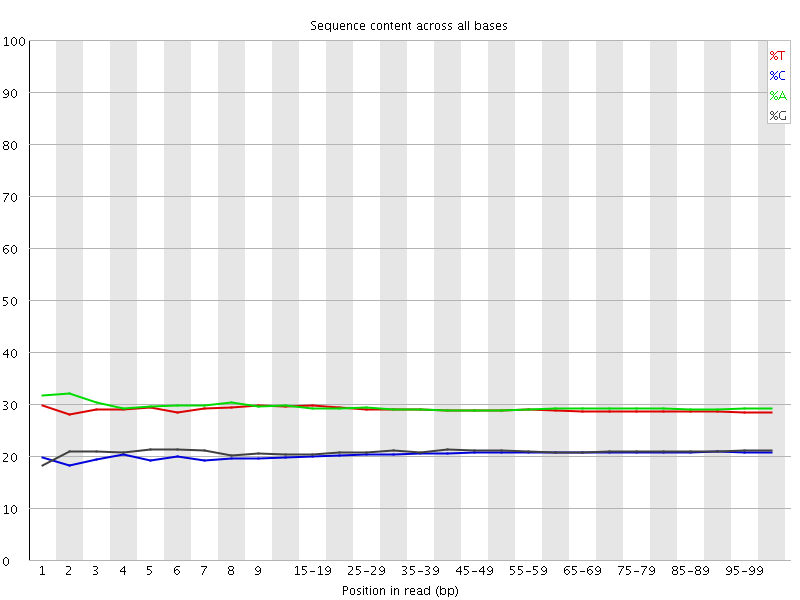
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
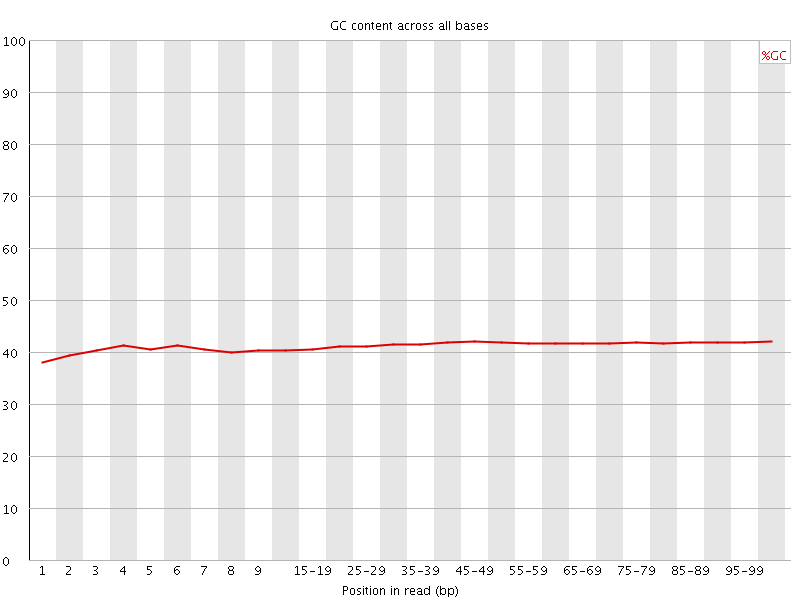
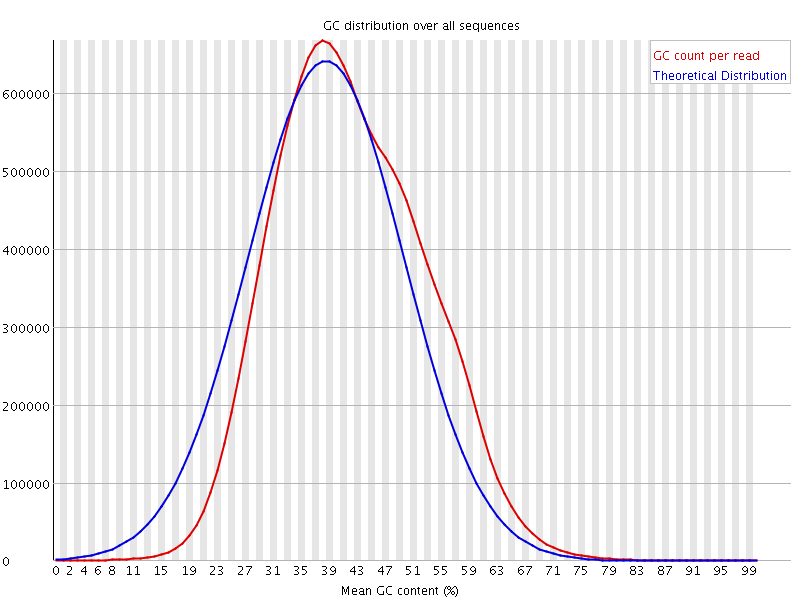
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
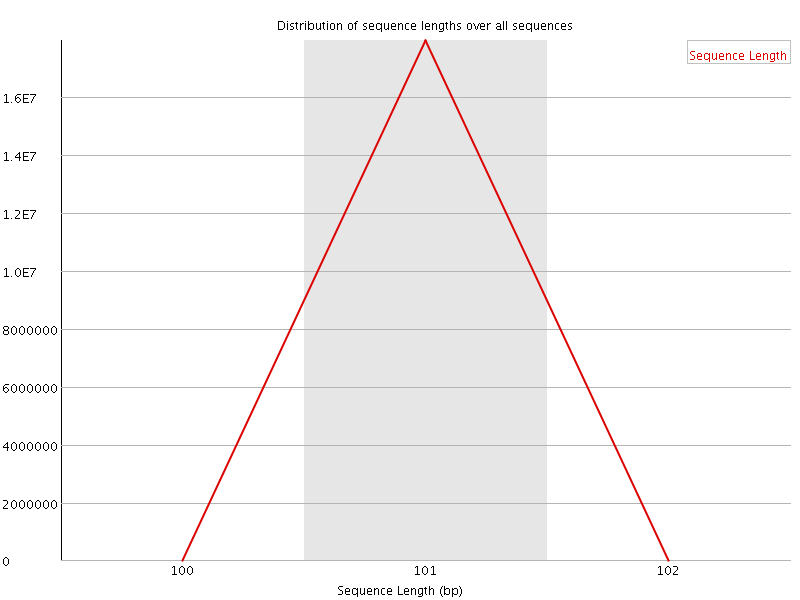
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
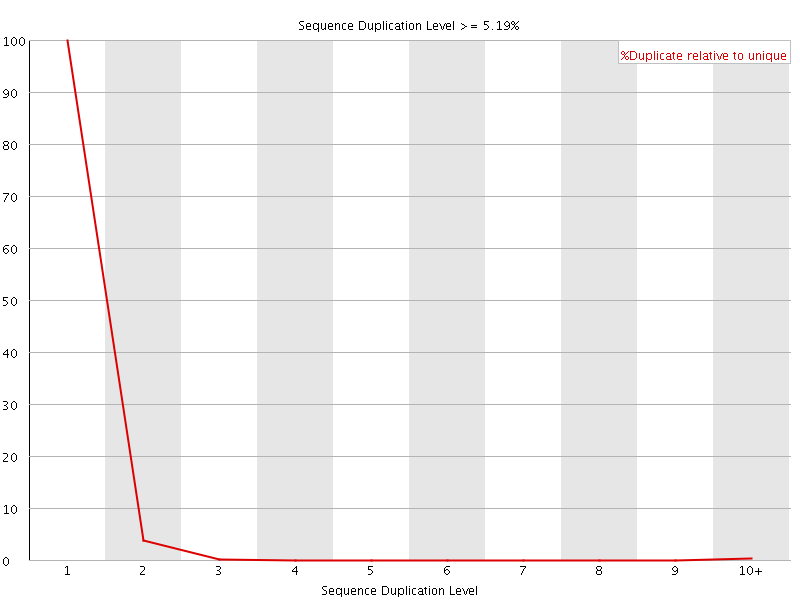
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
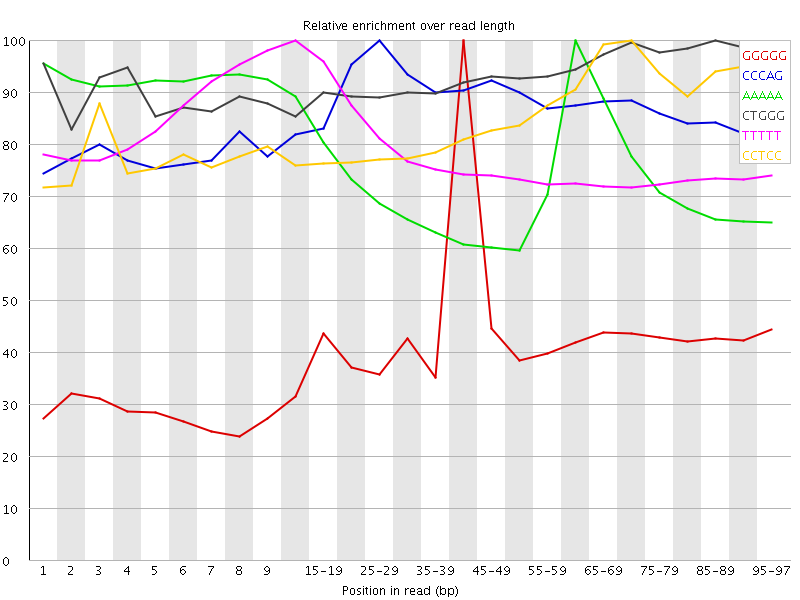
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2654975 | 3.749728 | 8.796886 | 40-44 |
| CCCAG | 3483685 | 3.719134 | 4.2561154 | 25-29 |
| AAAAA | 13776865 | 3.6407256 | 4.9242377 | 60-64 |
| CTGGG | 3363425 | 3.4940667 | 3.7450967 | 85-89 |
| TTTTT | 12070290 | 3.346834 | 4.2776804 | 10-14 |
| CCTCC | 3005215 | 3.2996583 | 3.8676074 | 70-74 |
| GGAGG | 3256405 | 3.289254 | 3.5523825 | 40-44 |
| CCAGG | 3131055 | 3.2815497 | 3.6932802 | 25-29 |
| CCTGG | 2948310 | 3.1198757 | 3.3976252 | 70-74 |
| GGGAG | 2910450 | 2.939809 | 5.4597244 | 5 |
| GGAAG | 2804435 | 2.025928 | 5.2582874 | 5 |
| AGAGC | 1902430 | 1.3999159 | 5.2870717 | 8 |
| GAGCG | 447495 | 0.46042815 | 5.9459968 | 9 |