![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL1_CGATGT_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 17934766 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
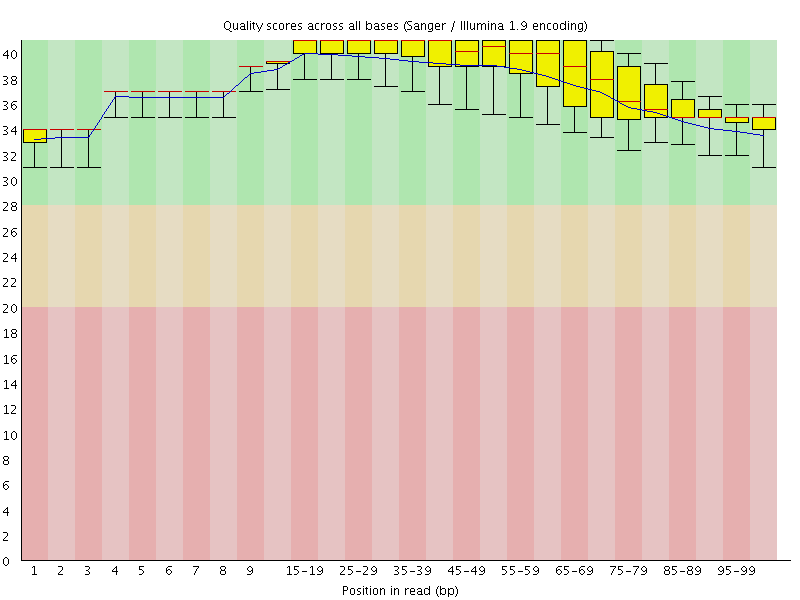
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
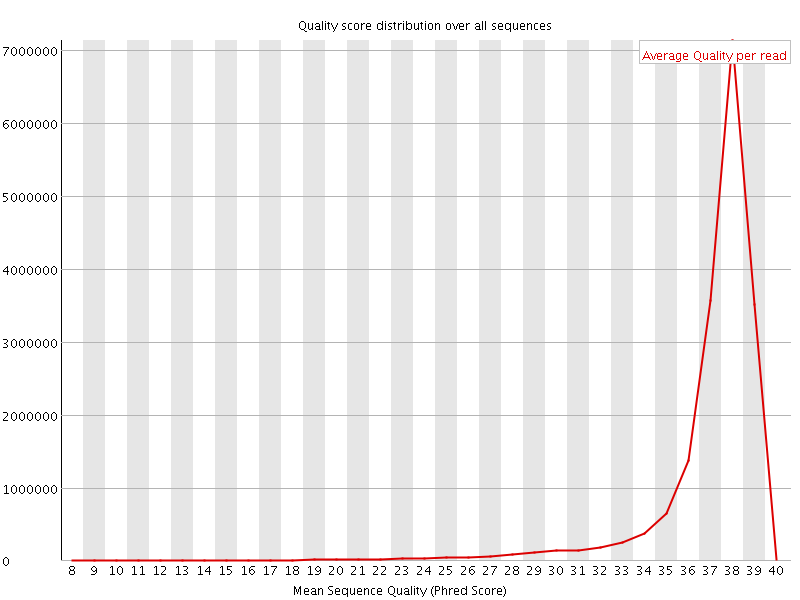
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
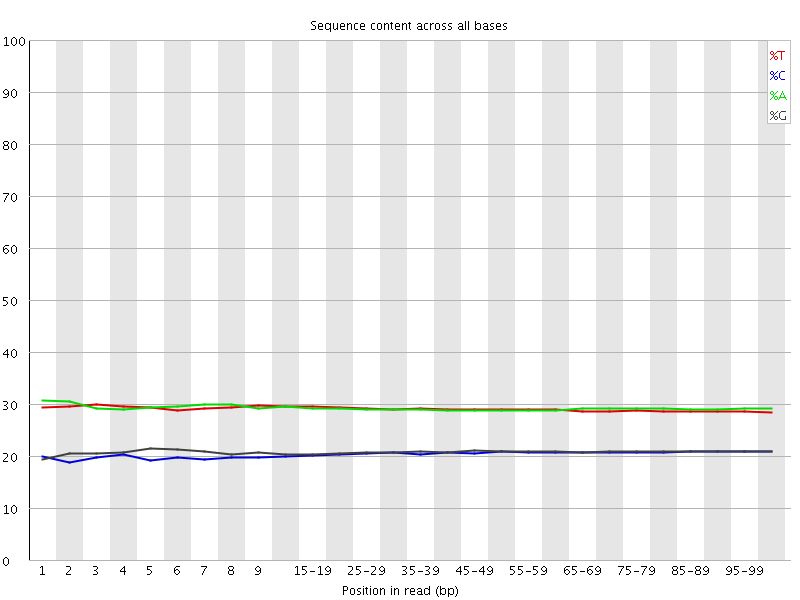
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
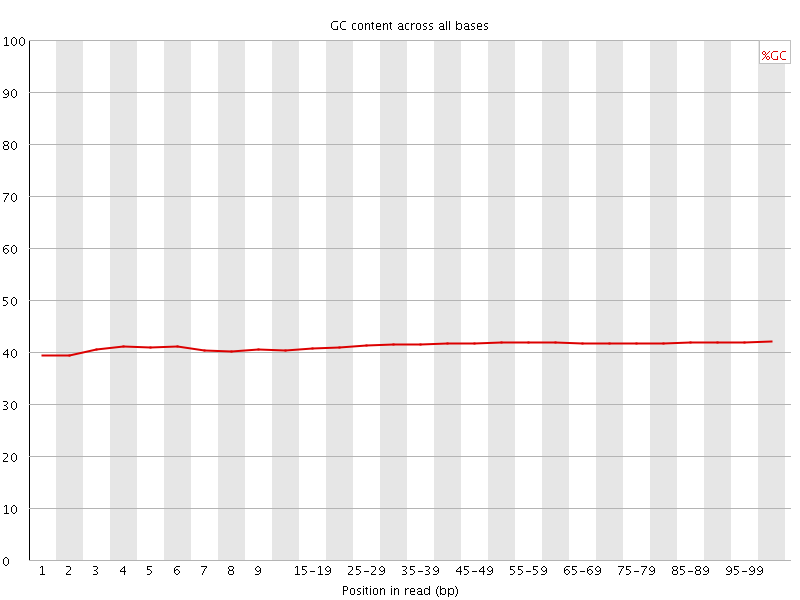
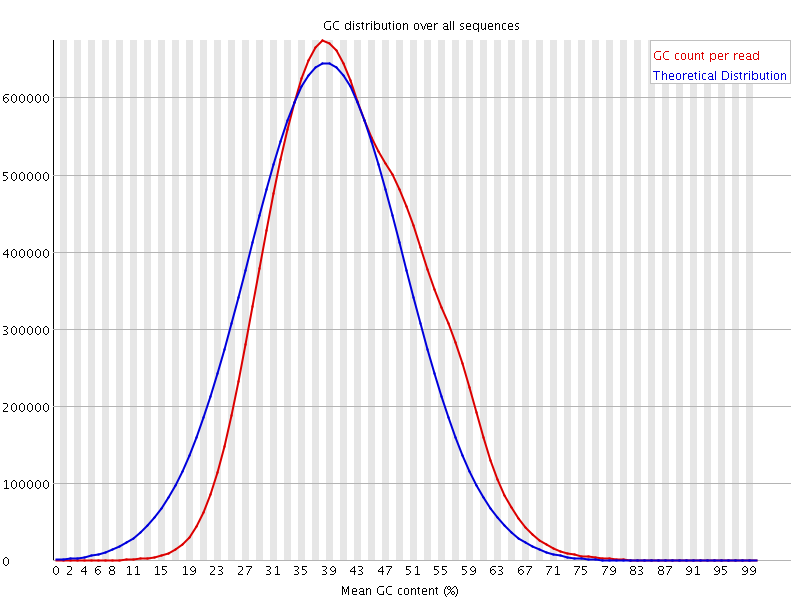
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
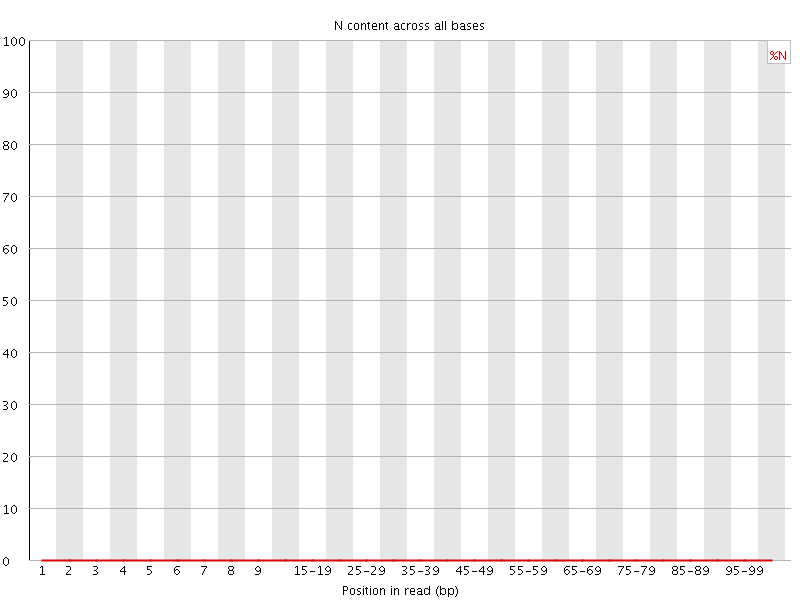
![[OK]](Icons/tick.png) Per base N content
Per base N content
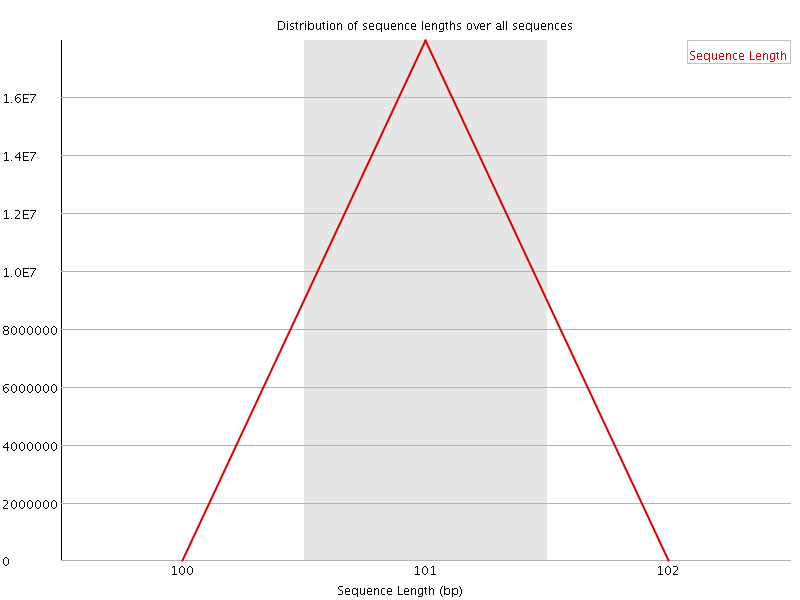
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
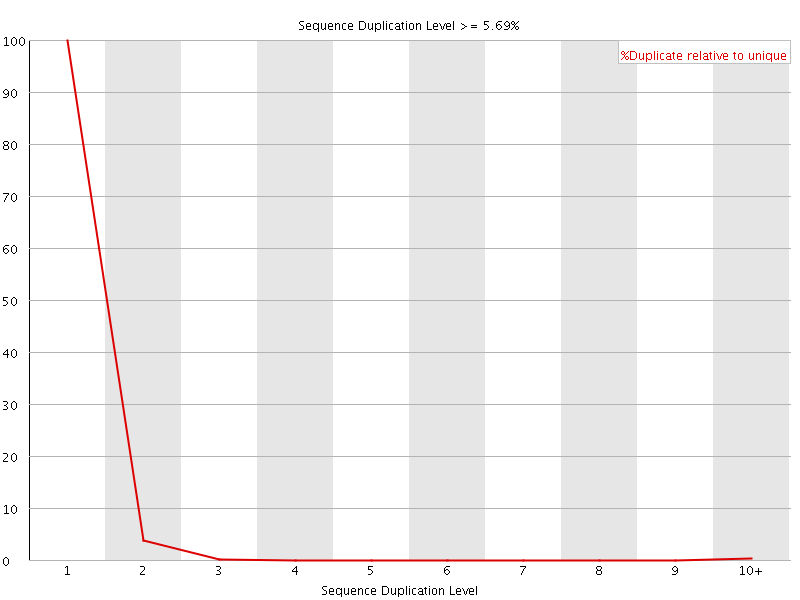
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGC | 91819 | 0.5119609589553608 | TruSeq Adapter, Index 2 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
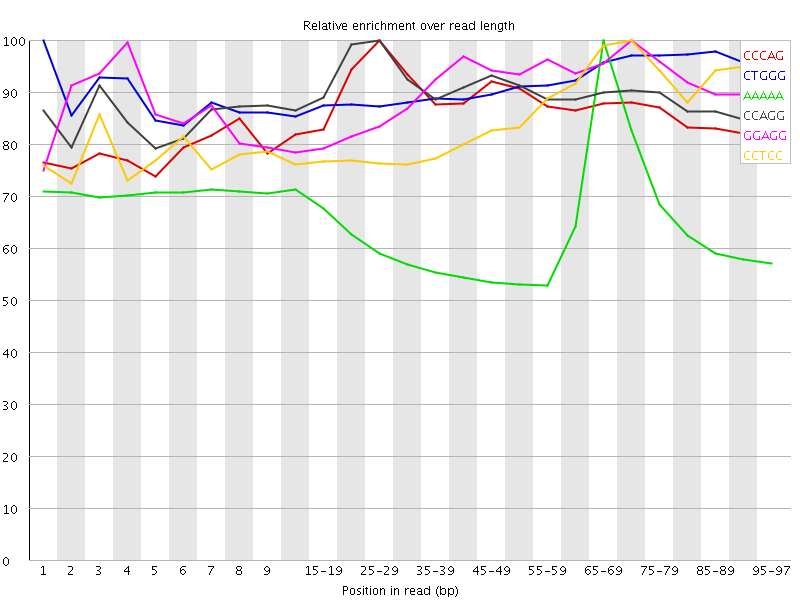
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCCAG | 3495735 | 3.7101305 | 4.2631497 | 25-29 |
| CTGGG | 3366480 | 3.518276 | 3.8398135 | 1 |
| AAAAA | 12636270 | 3.38564 | 5.275576 | 65-69 |
| CCAGG | 3136440 | 3.2966669 | 3.673093 | 25-29 |
| GGAGG | 3187400 | 3.2858624 | 3.6434844 | 70-74 |
| CCTCC | 3016825 | 3.24595 | 3.8054466 | 70-74 |
| TTTTT | 11737170 | 3.207949 | 3.844085 | 10-14 |
| CCTGG | 2964570 | 3.1284425 | 3.3790073 | 70-74 |
| GGAAG | 2796380 | 2.0583124 | 9.188917 | 5 |
| GAAGA | 2954230 | 1.5526075 | 6.683776 | 6 |
| AAGAG | 2949005 | 1.5498613 | 6.614726 | 7 |
| AGAGC | 1948750 | 1.4483844 | 8.512444 | 8 |
| GAGCA | 1881490 | 1.3983942 | 8.440017 | 9 |
| GATCG | 568775 | 0.4244208 | 7.5034513 | 1 |
| CGGAA | 534980 | 0.39761728 | 7.532784 | 4 |
| TCGGA | 473120 | 0.3530429 | 7.4982347 | 3 |
| ATCGG | 462255 | 0.34493542 | 7.486082 | 2 |